This is a preprint.
An assessment of the genomic structural variation landscape in Sub-Saharan African populations
- PMID: 39041024
- PMCID: PMC11261963
- DOI: 10.21203/rs.3.rs-4485126/v1
An assessment of the genomic structural variation landscape in Sub-Saharan African populations
Abstract
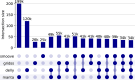
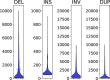
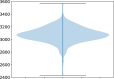
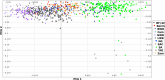
Structural variants are responsible for a large part of genomic variation between individuals and play a role in both common and rare diseases. Databases cataloguing structural variants notably do not represent the full spectrum of global diversity, particularly missing information from most African populations. To address this representation gap, we analysed 1,091 high-coverage African genomes, 545 of which are public data sets, and 546 which have been analysed for structural variants for the first time. Variants were called using five different tools and datasets merged and jointly called using SURVIVOR. We identified 67,795 structural variants throughout the genome, with 10,421 genes having at least one variant. Using a conservative overlap in merged data, 6,414 of the structural variants (9.5%) are novel compared to the Database of Genomic Variants. This study contributes to knowledge of the landscape of structural variant diversity in Africa and presents a reliable dataset for potential applications in population genetics and health-related research.
Keywords: African diversity; Structural variants; copy number variants; genomic variation.
Conflict of interest statement
Additional Declarations: There is NO Competing Interest.
Figures

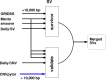
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

