Decoding the muscle transcriptome of patients with late-onset Pompe disease reveals markers of disease progression
- PMID: 39045638
- PMCID: PMC11629704
- DOI: 10.1093/brain/awae249
Decoding the muscle transcriptome of patients with late-onset Pompe disease reveals markers of disease progression
Abstract
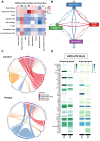
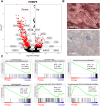
Late-onset Pompe disease (LOPD) is a rare genetic disorder caused by the deficiency of acid alpha-glucosidase leading to progressive cellular dysfunction owing to the accumulation of glycogen in the lysosome. The mechanism of relentless muscle damage (a classic manifestation of the disease) has been studied extensively by analysing the whole-muscle tissue; however, little, if anything, is known about transcriptional heterogeneity among nuclei within the multinucleated skeletal muscle cells. This is the first report of application of single-nucleus RNA sequencing to uncover changes in the gene expression profile in muscle biopsies from eight patients with LOPD and four muscle samples from age- and sex-matched healthy controls. We matched these changes with histological findings using GeoMx spatial transcriptomics to compare the transcriptome of control myofibres from healthy individuals with non-vacuolated (histologically unaffected) and vacuolated (histologically affected) myofibres of LODP patients. We observed an increase in the proportion of slow and regenerative muscle fibres and macrophages in LOPD muscles. The expression of the genes involved in glycolysis was reduced, whereas the expression of the genes involved in the metabolism of lipids and amino acids was increased in non-vacuolated fibres, indicating early metabolic abnormalities. Additionally, we detected upregulation of autophagy genes and downregulation of the genes involved in ribosomal and mitochondrial function leading to defective oxidative phosphorylation. Upregulation of genes associated with inflammation, apoptosis and muscle regeneration was observed only in vacuolated fibres. Notably, enzyme replacement therapy (the only available therapy for the disease) showed a tendency to restore dysregulated metabolism, particularly within slow fibres. A combination of single-nucleus RNA sequencing and spatial transcriptomics revealed the landscape of the normal and diseased muscle and highlighted the early abnormalities associated with disease progression. Thus, the application of these two new cutting-edge technologies provided insight into the molecular pathophysiology of muscle damage in LOPD and identified potential avenues for therapeutic intervention.
Keywords: glycogen storage disease; metabolism; mitochondria abnormality.
© The Author(s) 2024. Published by Oxford University Press on behalf of the Guarantors of Brain.
Conflict of interest statement
The authors report no competing interests.
Figures




References
-
- van der Ploeg AT, Reuser AJ. Pompe’s disease. Lancet. 2008;372:1342–1353. - PubMed
-
- Case LE, Beckemeyer AA, Kishnani PS. Infantile Pompe disease on ERT—Update on clinical presentation, musculoskeletal management, and exercise considerations. Am J Med Genet C Semin Med Genet. 2012;160C:69–79. - PubMed
-
- Schänzer A, Kaiser AK, Mühlfeld C, et al. . Quantification of muscle pathology in infantile Pompe disease. Neuromuscul Disord. 2017;27:141–152. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

