Seeing in the dark: a metagenomic approach can illuminate the drivers of plant disease
- PMID: 39055364
- PMCID: PMC11269093
- DOI: 10.3389/fpls.2024.1405042
Seeing in the dark: a metagenomic approach can illuminate the drivers of plant disease
Keywords: functional metagenomics; metagenomics; microbiome; multi-omics; novel genes; plant diagnostics; plant pathogen; workflow.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
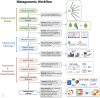

References
-
- Agrios G. (2005). Plant pathology., 5th edn (Amsterdam: Elsevier Academic Press; ).
-
- Aravind R., Kumar A., Eapen S. J., Ramana K. V. (2009). Endophytic bacterial flora in root and stem tissues of black pepper (Piper nigrum L.) genotype: isolation, identification and evaluation against Phytophthora capsici. Lett. Appl. Microbiol. 48, 58–64. doi: 10.1111/lam.2009.48.issue-1 - DOI - PubMed
LinkOut - more resources
Full Text Sources

