Chromosomal structural rearrangements implicate long non-coding RNAs in rare germline disorders
- PMID: 39060644
- PMCID: PMC11294402
- DOI: 10.1007/s00439-024-02693-y
Chromosomal structural rearrangements implicate long non-coding RNAs in rare germline disorders
Abstract
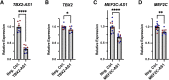
In recent years, there has been increased focus on exploring the role the non-protein-coding genome plays in Mendelian disorders. One class of particular interest is long non-coding RNAs (lncRNAs), which has recently been implicated in the regulation of diverse molecular processes. However, because lncRNAs do not encode protein, there is uncertainty regarding what constitutes a pathogenic lncRNA variant, and thus annotating such elements is challenging. The Developmental Genome Anatomy Project (DGAP) and similar projects recruit individuals with apparently balanced chromosomal abnormalities (BCAs) that disrupt or dysregulate genes in order to annotate the human genome. We hypothesized that rearrangements disrupting lncRNAs could be the underlying genetic etiology for the phenotypes of a subset of these individuals. Thus, we assessed 279 cases with BCAs and selected 191 cases with simple BCAs (breakpoints at only two genomic locations) for further analysis of lncRNA disruptions. From these, we identified 66 cases in which the chromosomal rearrangements directly disrupt lncRNAs. In 30 cases, no genes of any other class aside from lncRNAs are directly disrupted, consistent with the hypothesis that lncRNA disruptions could underly the phenotypes of these individuals. Strikingly, the lncRNAs MEF2C-AS1 and ENSG00000257522 are each disrupted in two unrelated cases. Furthermore, we experimentally tested the lncRNAs TBX2-AS1 and MEF2C-AS1 and found that knockdown of these lncRNAs resulted in decreased expression of the neighboring transcription factors TBX2 and MEF2C, respectively. To showcase the power of this genomic approach for annotating lncRNAs, here we focus on clinical reports and genetic analysis of seven individuals with likely developmental etiologies due to lncRNA disruptions.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures







Update of
-
Rare germline disorders implicate long non-coding RNAs disrupted by chromosomal structural rearrangements.medRxiv [Preprint]. 2024 Jun 19:2024.06.16.24307499. doi: 10.1101/2024.06.16.24307499. medRxiv. 2024. Update in: Hum Genet. 2024 Jul;143(7):921-938. doi: 10.1007/s00439-024-02693-y. PMID: 38946951 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

