The transcriptional regulator Fur modulates the expression of uge, a gene essential for the core lipopolysaccharide biosynthesis in Klebsiella pneumoniae
- PMID: 39061004
- PMCID: PMC11282780
- DOI: 10.1186/s12866-024-03418-x
The transcriptional regulator Fur modulates the expression of uge, a gene essential for the core lipopolysaccharide biosynthesis in Klebsiella pneumoniae
Abstract
Background: Klebsiella pneumoniae is a Gram-negative pathogen that has become a threat to public health worldwide due to the emergence of hypervirulent and multidrug-resistant strains. Cell-surface components, such as polysaccharide capsules, fimbriae, and lipopolysaccharides (LPS), are among the major virulence factors for K. pneumoniae. One of the genes involved in LPS biosynthesis is the uge gene, which encodes the uridine diphosphate galacturonate 4-epimerase enzyme. Although essential for the LPS formation in K. pneumoniae, little is known about the mechanisms that regulate the expression of uge. Ferric uptake regulator (Fur) is an iron-responsive transcription factor that modulates the expression of capsular and fimbrial genes, but its role in LPS expression has not yet been identified. This work aimed to investigate the role of the Fur regulator in the expression of the K. pneumoniae uge gene and to determine whether the production of LPS by K. pneumoniae is modulated by the iron levels available to the bacterium.
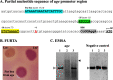
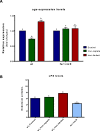
Results: Using bioinformatic analyses, a Fur-binding site was identified on the promoter region of the uge gene; this binding site was validated experimentally through Fur Titration Assay (FURTA) and DNA Electrophoretic Mobility Shift Assay (EMSA) techniques. RT-qPCR analyses were used to evaluate the expression of uge according to the iron levels available to the bacterium. The iron-rich condition led to a down-regulation of uge, while the iron-restricted condition resulted in up-regulation. In addition, LPS was extracted and quantified on K. pneumoniae cells subjected to iron-replete and iron-limited conditions. The iron-limited condition increased the amount of LPS produced by K. pneumoniae. Finally, the expression levels of uge and the amount of the LPS were evaluated on a K. pneumoniae strain mutant for the fur gene. Compared to the wild-type, the strain with the fur gene knocked out presented a lower LPS amount and an unchanged expression of uge, regardless of the iron levels.
Conclusions: Here, we show that iron deprivation led the K. pneumoniae cells to produce higher amount of LPS and that the Fur regulator modulates the expression of uge, a gene essential for LPS biosynthesis. Thus, our results indicate that iron availability modulates the LPS biosynthesis in K. pneumoniae through a Fur-dependent mechanism.
Keywords: Klebsiella pneumoniae; Gene expression analyses; Iron-regulatory proteins; Lipopolysaccharides; Transcriptional regulation.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

