Protein Assemblies in Translesion Synthesis
- PMID: 39062611
- PMCID: PMC11276120
- DOI: 10.3390/genes15070832
Protein Assemblies in Translesion Synthesis
Abstract
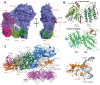
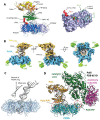
Translesion synthesis (TLS) is a mechanism of DNA damage tolerance utilized by eukaryotic cells to replicate DNA across lesions that impede the high-fidelity replication machinery. In TLS, a series of specialized DNA polymerases are employed, which recognize specific DNA lesions, insert nucleotides across the damage, and extend the distorted primer-template. This allows cells to preserve genetic integrity at the cost of mutations. In humans, TLS enzymes include the Y-family, inserter polymerases, Polη, Polι, Polκ, Rev1, and the B-family extender polymerase Polζ, while in S. cerevisiae only Polη, Rev1, and Polζ are present. To bypass DNA lesions, TLS polymerases cooperate, assembling into a complex on the eukaryotic sliding clamp, PCNA, termed the TLS mutasome. The mutasome assembly is contingent on protein-protein interactions (PPIs) between the modular domains and subunits of TLS enzymes, and their interactions with PCNA and DNA. While the structural mechanisms of DNA lesion bypass by the TLS polymerases and PPIs of their individual modules are well understood, the mechanisms by which they cooperate in the context of TLS complexes have remained elusive. This review focuses on structural studies of TLS polymerases and describes the case of TLS holoenzyme assemblies in action emerging from recent high-resolution Cryo-EM studies.
Keywords: DNA damage tolerance; DNA repair; protein assemblies; protein structure; protein–protein interactions; translesion synthesis.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures




Similar articles
-
Interaction between the Rev1 C-Terminal Domain and the PolD3 Subunit of Polζ Suggests a Mechanism of Polymerase Exchange upon Rev1/Polζ-Dependent Translesion Synthesis.Biochemistry. 2016 Apr 5;55(13):2043-53. doi: 10.1021/acs.biochem.5b01282. Epub 2016 Mar 24. Biochemistry. 2016. PMID: 26982350 Free PMC article.
-
NMR mapping of PCNA interaction with translesion synthesis DNA polymerase Rev1 mediated by Rev1-BRCT domain.J Mol Biol. 2013 Sep 9;425(17):3091-105. doi: 10.1016/j.jmb.2013.05.029. Epub 2013 Jun 7. J Mol Biol. 2013. PMID: 23747975
-
Structural basis for novel interactions between human translesion synthesis polymerases and proliferating cell nuclear antigen.J Biol Chem. 2009 Apr 17;284(16):10552-60. doi: 10.1074/jbc.M809745200. Epub 2009 Feb 10. J Biol Chem. 2009. PMID: 19208623 Free PMC article.
-
The Rev1-Polζ translesion synthesis mutasome: Structure, interactions and inhibition.Enzymes. 2019;45:139-181. doi: 10.1016/bs.enz.2019.07.001. Epub 2019 Aug 9. Enzymes. 2019. PMID: 31627876 Free PMC article. Review.
-
Ubiquitin-dependent regulation of translesion polymerases.Biochem Soc Trans. 2010 Feb;38(Pt 1):110-5. doi: 10.1042/BST0380110. Biochem Soc Trans. 2010. PMID: 20074045 Review.
Cited by
-
Targeting the DNA damage response in cancer.MedComm (2020). 2024 Oct 31;5(11):e788. doi: 10.1002/mco2.788. eCollection 2024 Nov. MedComm (2020). 2024. PMID: 39492835 Free PMC article. Review.
References
-
- Friedberg E.C., Wood R.D., Walker G.C., Schultz R.A., Slede W., Ellenberger T. DNA Repair and Mutagenesis. 2nd ed. ASM Press; Washington, DC, USA: 2006.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

