Genome-Wide Characterization and Expression Profiling of the AP2/ERF Gene Family in Fragaria vesca L
- PMID: 39062854
- PMCID: PMC11277216
- DOI: 10.3390/ijms25147614
Genome-Wide Characterization and Expression Profiling of the AP2/ERF Gene Family in Fragaria vesca L
Abstract
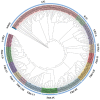
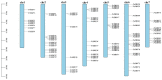
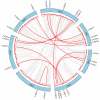
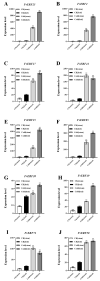
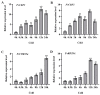
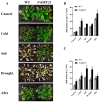
The wild strawberry (Fragaria vesca L.; F. vesca) represents a resilient and extensively studied model organism. While the AP2/ERF gene family plays a pivotal role in plant development, its exploration within F. vesca remains limited. In this study, we characterized the AP2/ERF gene family in wild strawberries using the recently released genomic data (F. vesca V6.0). We conducted an analysis of the gene family expansion pattern, we examined gene expression in stem segments and leaves under cold conditions, and we explored its functional attributes. Our investigation revealed that the FvAP2/ERF family comprises 86 genes distributed among four subfamilies: AP2 (17), RAV (6), ERF (62), and Soloist (1). Tandem and segmental duplications significantly contributed to the growth of this gene family. Furthermore, predictive analysis identified several cis-acting elements in the promoter region associated with meristematic tissue expression, hormone regulation, and resistance modulation. Transcriptomic analysis under cold stress unveiled diverse responses among multiple FvAP2/ERFs in stem segments and leaves. Real-time fluorescence quantitative reverse transcription PCR (RT-qPCR) results confirmed elevated expression levels of select genes following the cold treatment. Additionally, overexpression of FvERF23 in Arabidopsis enhanced cold tolerance, resulting in significantly increased fresh weight and root length compared to the wild-type control. These findings lay the foundation for further exploration into the functional roles of FvAP2/ERF genes.
Keywords: AP2/ERF; Fragaria vesca L.; abiotic stresses; cold; transcription factor.
Conflict of interest statement
The authors declare that they have no conflicts of interest regarding this research.
Figures



References
-
- Jan N., Hussain M.U., Andrabi K.I. Cold resistance in plants: A mystery unresolved. Electron. J. Biotechnol. 2009;12:14–15. doi: 10.2225/vol12-issue3-fulltext-3. - DOI
MeSH terms
Substances
Grants and funding
- 32172521/National Natural Science Foundation of China
- LH2022C023/Natural Science Foundation of Heilongjiang Province, China
- 2023MD744175/the China Postdoctoral Science Foundation
- 2022YFD1600501-13/the National Key Research and Development Program of China
- No/the Modern Agricultural Industrial Technology Collaborative Innovation and Promotion System of Heilongjiang Province
LinkOut - more resources
Full Text Sources
Research Materials

