Transcriptome Analysis of Mesenchymal Progenitor Cells Revealed Molecular Insights into Metabolic Dysfunction and Inflammation in Polycystic Ovary Syndrome
- PMID: 39063189
- PMCID: PMC11276887
- DOI: 10.3390/ijms25147948
Transcriptome Analysis of Mesenchymal Progenitor Cells Revealed Molecular Insights into Metabolic Dysfunction and Inflammation in Polycystic Ovary Syndrome
Abstract
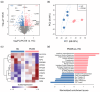
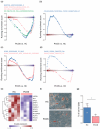
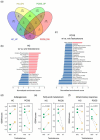
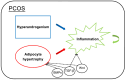
Polycystic ovary syndrome (PCOS) is a female endocrine disorder with metabolic issues. Hyperandrogenism combined with hyperinsulinemia exacerbates the reproductive, metabolic, and inflammatory problems in PCOS patients. The etiology of PCOS is unclear. Patient-specific induced pluripotent stem cells (iPSCs) offer a promising model for studying disease mechanisms and conducting drug screening. Here, we aim to use mesenchymal progenitor cells (MPCs) derived from PCOS iPSCs to explore the mechanism of PCOS. We compared the transcriptome profiles of PCOS and healthy control (HC) iPSC-derived MPCs (iPSCMs). Moreover, we assess the impact of androgens on iPSCMs. In the comparison between PCOS and HC, the expression levels of 1026 genes were significantly different. A gene set enrichment analysis (GSEA) revealed that adipogenesis- and metabolism-related genes were downregulated, whereas inflammation-related genes were upregulated in the PCOS iPSCMs. Dysregulation of the TGF-β1 and Wnt signaling pathways was observed in the PCOS iPSCMs. Furthermore, there was impaired adipogenesis and decreased lipolysis in the PCOS iPSCMs-derived adipocytes. With testosterone treatment, genes related to metabolism were upregulated in the HC iPSCMs but downregulated in the PCOS iPSCMs. The impact of testosterone varied among HCs and PCOS iPSCMs, possibly because of a genetic predisposition toward PCOS. This study found specific signaling pathways that could serve as therapeutic targets for PCOS.
Keywords: RNA sequencing; induced pluripotent stem cells; polycystic ovary syndrome.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures



References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

