Disease-associated gut microbiome and metabolome changes in rats with chronic hypoxia-induced pulmonary hypertension
- PMID: 39071798
- PMCID: PMC11272533
- DOI: 10.3389/fcell.2024.1022181
Disease-associated gut microbiome and metabolome changes in rats with chronic hypoxia-induced pulmonary hypertension
Abstract
Background: Pulmonary hypertension (PH) is a progressive disease affecting the lung vasculature that is characterized by sustained vasoconstriction and leads to vascular remodeling. The lung microbiome contributes to PH progression, but the function of the gut microbiome and the correlation between the gut microbiome and metabolome remain unclear. We have analyzed whether chronic hypoxia-induced PH alters the rat fecal microbiota.
Purpose: We explored hypoxia-induced pulmonary hypertension model rats to find out the characteristic changes of intestinal microorganisms and metabolites of hypoxia-induced pulmonary hypertension, and provide a theoretical basis for clinical treatment.
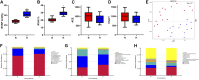
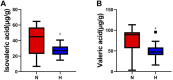
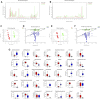
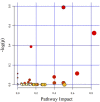
Methods: In the current study, a chronic hypoxia-induced PH rat model was used to investigate the role of the gut microbiome and metabolome as a potential mechanism contributing to the occurrence and development of PH. 16S ribosomal ribonucleic acid (16S rRNA), short-chain fatty acid (SCFA) measurements, mass spectrometry (MS) metabolomics analysis and metatranscriptome were performed to analyze stool samples. The datasets were analyzed individually and integrated for combined analysis using bioinformatics approaches.
Results: Our results suggest that the gut microbiome and metabolome of chronic hypoxia-induced PH rats are distinct from those of normoxic rats and may thus aid in the search for new therapeutic or diagnostic paradigms for PH.
Conclusion: The gut microbiome and metabolome are altered as a result of chronic hypoxia-induced PH. This imbalanced bacterial ecosystem might play a pathophysiological role in PH by altering homeostasis.
Keywords: SCFAs; chronic hypoxia; gut metabolome; gut microbiome; pH.
Copyright © 2024 Cao, Wang, Mo, Peng, Hong, Zhou, Sun, Li, Liang, Zhao, Zheng, Li and Peng.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

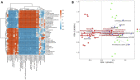


References
-
- Boronat A., Mateus J., Soldevila-Domenech N., Guerra M., Rodríguez-Morató J., Varon C., et al. (2019). Cardiovascular benefits of tyrosol and its endogenous conversion into hydroxytyrosol in humans. A randomized, controlled trial. Free Radic. Biol. Med. 143, 471–481. 10.1016/j.freeradbiomed.2019.08.032 - DOI - PubMed
LinkOut - more resources
Full Text Sources

