Engineering Toxoplasma gondii secretion systems for intracellular delivery of multiple large therapeutic proteins to neurons
- PMID: 39075233
- PMCID: PMC11306108
- DOI: 10.1038/s41564-024-01750-6
Engineering Toxoplasma gondii secretion systems for intracellular delivery of multiple large therapeutic proteins to neurons
Abstract
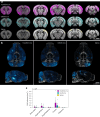
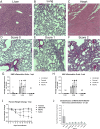
Delivering macromolecules across biological barriers such as the blood-brain barrier limits their application in vivo. Previous work has demonstrated that Toxoplasma gondii, a parasite that naturally travels from the human gut to the central nervous system (CNS), can deliver proteins to host cells. Here we engineered T. gondii's endogenous secretion systems, the rhoptries and dense granules, to deliver multiple large (>100 kDa) therapeutic proteins into neurons via translational fusions to toxofilin and GRA16. We demonstrate delivery in cultured cells, brain organoids and in vivo, and probe protein activity using imaging, pull-down assays, scRNA-seq and fluorescent reporters. We demonstrate robust delivery after intraperitoneal administration in mice and characterize 3D distribution throughout the brain. As proof of concept, we demonstrate GRA16-mediated brain delivery of the MeCP2 protein, a putative therapeutic target for Rett syndrome. By characterizing the potential and current limitations of the system, we aim to guide future improvements that will be required for broader application.
© 2024. The Author(s).
Conflict of interest statement
S.B., O.R. and L.S. are inventors on a patent application filed by Tel-Aviv University and The University of Glasgow related to this work (PCT/IL2017/050731, WO2018002938A1), and are scientific advisors of Epeius Pharma Ltd. N.M., H.L., R.E., V.K. and D.L. are employees of Epeius Pharma Ltd. The remaining authors declare no competing interests.
Figures









Comment in
-
Engineered parasite delivers proteins to the brain.Nat Rev Drug Discov. 2024 Oct;23(10):740. doi: 10.1038/d41573-024-00137-w. Nat Rev Drug Discov. 2024. PMID: 39191964 No abstract available.
References
-
- Solaro, R., Chiellini, F. & Battisti, A. Targeted delivery of protein drugs by nanocarriers. Materials3, 1928–1980 (2010).10.3390/ma3031928 - DOI
MeSH terms
Substances
Grants and funding
- P30 CA023074/CA/NCI NIH HHS/United States
- 104111/WT_/Wellcome Trust/United Kingdom
- P30CA023074/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
- WT_/Wellcome Trust/United Kingdom
- R01 NS095994/NS/NINDS NIH HHS/United States
- Basic Research grant/International Rett Syndrome Foundation (IRSF)
- Personal Research Fellowship/Royal Society of Edinburgh (RSE)
- T32 AG061897/AG/NIA NIH HHS/United States
- 341117/EC | EU Framework Programme for Research and Innovation H2020 | H2020 Priority Excellent Science | H2020 European Research Council (H2020 Excellent Science - European Research Council)
- T32AG061897/U.S. Department of Health & Human Services | NIH | National Institute on Aging (U.S. National Institute on Aging)
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

