Protocol for calculating binding free energy of RNA:RNA interactions through molecular dynamics simulations using adaptive biasing force technique
- PMID: 39083381
- PMCID: PMC11342170
- DOI: 10.1016/j.xpro.2024.103223
Protocol for calculating binding free energy of RNA:RNA interactions through molecular dynamics simulations using adaptive biasing force technique
Abstract
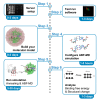
The adaptive biasing force (ABF) technique allows sampling to proceed in a flat free energy surface when performing molecular dynamics (MD) simulations. Here, we present a protocol to perform MD simulations using the ABF technique and apply it to calculate the binding free energy of an RNA:RNA interaction. We describe steps for server setup, test running software, and building molecular models. We then detail procedures for running and configuring ABF-MD simulations and analyzing binding free energy and structural change. For complete details on the use and execution of this protocol, please refer to Fujita et al.1 and Kameda et al.2.
Keywords: Biophysics; Biotechnology and bioengineering; Computer sciences.
Copyright © 2024 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures

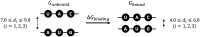
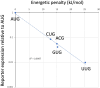
References
-
- Moore P.B., Steitz T.A. The involvement of RNA in ribosome function. Nature. 2002;418:229–235. - PubMed
-
- Doudna J.A., Lorsch J.R. Ribozyme catalysis: not different, just worse. Nat. Struct. Mol. Biol. 2005;12:395–402. - PubMed
-
- Asano K. In: Translational Control in Encyclopedia of Systems Biology. Werner D., Wolkenhauser O., Cho K.-H., Yokota H., editors. Springer; 2013. pp. 2278–2282.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

