ATM and 53BP1 regulate alternative end joining-mediated V(D)J recombination
- PMID: 39083600
- PMCID: PMC11290492
- DOI: 10.1126/sciadv.adn4682
ATM and 53BP1 regulate alternative end joining-mediated V(D)J recombination
Abstract
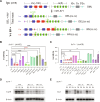
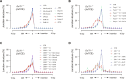
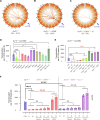
G0-G1 phase alternative end joining (A-EJ) is a recently defined mutagenic pathway characterized by resected deletion and translocation joints that are predominantly direct and are distinguished from A-EJ in cycling cells that rely much more on microhomology-mediated end joining (MMEJ). Using chemical and genetic approaches, we systematically evaluate potential A-EJ factors and DNA damage response (DDR) genes to support this mechanism by mapping the repair fates of RAG1/2-initiated double-strand breaks in the context of Igκ locus V-J recombination and chromosome translocation. Our findings highlight a polymerase theta-independent Parp1-XRCC1/LigIII axis as central A-EJ components, supported by 53BP1 in the context of an Ataxia-telangiectasia mutated (ATM)-activated DDR. Mechanistically, we demonstrate varied changes in short-range resection, MMEJ, and translocation, imposed by compromising specific DDR activities, which include polymerase alpha, Ataxia-telangiectasia and Rad3-related (ATR), DNA2, and Mre11. This study advances our understanding of DNA damage repair within the 53BP1 regulatory domain and the RAG1/2 postcleavage complex.
Figures





References
-
- Schatz D. G., Ji Y., Recombination centres and the orchestration of V(D)J recombination. Nat. Rev. Immunol. 11, 251–263 (2011). - PubMed
-
- Liu Q., Palomero L., Moore J., Guix I., Espín R., Aytés A., Mao J.-H., Paulovich A. G., Whiteaker J. R., Ivey R. G., Iliakis G., Luo D., Chalmers A. J., Murnane J., Pujana M. A., Barcellos-Hoff M. H., Loss of TGFβ signaling increases alternative end-joining DNA repair that sensitizes to genotoxic therapies across cancer types. Sci. Transl. Med. 13, eabc4465 (2021). - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

