The fork protection complex promotes parental histone recycling and epigenetic memory
- PMID: 39094569
- PMCID: PMC11383432
- DOI: 10.1016/j.cell.2024.07.017
The fork protection complex promotes parental histone recycling and epigenetic memory
Abstract
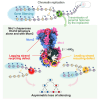
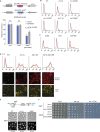
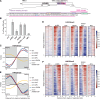
The inheritance of parental histones across the replication fork is thought to mediate epigenetic memory. Here, we reveal that fission yeast Mrc1 (CLASPIN in humans) binds H3-H4 tetramers and operates as a central coordinator of symmetric parental histone inheritance. Mrc1 mutants in a key connector domain disrupted segregation of parental histones to the lagging strand comparable to Mcm2 histone-binding mutants. Both mutants showed clonal and asymmetric loss of H3K9me-mediated gene silencing. AlphaFold predicted co-chaperoning of H3-H4 tetramers by Mrc1 and Mcm2, with the Mrc1 connector domain bridging histone and Mcm2 binding. Biochemical and functional analysis validated this model and revealed a duality in Mrc1 function: disabling histone binding in the connector domain disrupted lagging-strand recycling while another histone-binding mutation impaired leading strand recycling. We propose that Mrc1 toggles histones between the lagging and leading strand recycling pathways, in part by intra-replisome co-chaperoning, to ensure epigenetic transmission to both daughter cells.
Keywords: Claspin; DNA replication; H3K9 methylation; chromatin replication; epigenetic inheritance; epigenome maintenance; fission yeast; heterochromatin; histone chaperone; histone recycling; mouse embryonic stem cells.
Copyright © 2024 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests A.G. is co-founder and chief scientific officer (CSO) of Ankrin Therapeutics. A.G. is a member of the scientific advisory board of Molecular Cell. G.M. is a stockholder of Ensoma and a member of its scientific advisory board. The other authors declare no competing interests.
Figures












References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

