Development of an epigenetic clock resistant to changes in immune cell composition
- PMID: 39095531
- PMCID: PMC11297166
- DOI: 10.1038/s42003-024-06609-4
Development of an epigenetic clock resistant to changes in immune cell composition
Abstract
Epigenetic clocks are age predictors that use machine-learning models trained on DNA CpG methylation values to predict chronological or biological age. Increases in predicted epigenetic age relative to chronological age (epigenetic age acceleration) are connected to aging-associated pathologies, and changes in epigenetic age are linked to canonical aging hallmarks. However, epigenetic clocks rely on training data from bulk tissues whose cellular composition changes with age. Here, we found that human naive CD8+ T cells, which decrease in frequency during aging, exhibit an epigenetic age 15-20 years younger than effector memory CD8+ T cells from the same individual. Importantly, homogenous naive T cells isolated from individuals of different ages show a progressive increase in epigenetic age, indicating that current epigenetic clocks measure two independent variables, aging and immune cell composition. To isolate the age-associated cell intrinsic changes, we created an epigenetic clock, the IntrinClock, that did not change among 10 immune cell types tested. IntrinClock shows a robust predicted epigenetic age increase in a model of replicative senescence in vitro and age reversal during OSKM-mediated reprogramming.
© 2024. The Author(s).
Conflict of interest statement
A.T. and E.V. are listed co-inventors on pending patents relating to work disclosed in this manuscript. Remaining authors declare no competing interests.
Figures

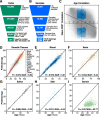



References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

