Regulatory role of lncH19 in RAC1 alternative splicing: implication for RAC1B expression in colorectal cancer
- PMID: 39098911
- PMCID: PMC11299361
- DOI: 10.1186/s13046-024-03139-z
Regulatory role of lncH19 in RAC1 alternative splicing: implication for RAC1B expression in colorectal cancer
Abstract
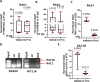
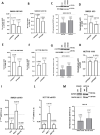
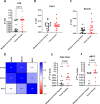
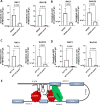
Aberrant alternative splicing events play a critical role in cancer biology, contributing to tumor invasion, metastasis, epithelial-mesenchymal transition, and drug resistance. Recent studies have shown that alternative splicing is a key feature for transcriptomic variations in colorectal cancer, which ranks third among malignant tumors worldwide in both incidence and mortality. Long non-coding RNAs can modulate this process by acting as trans-regulatory agents, recruiting splicing factors, or driving them to specific targeted genes. LncH19 is a lncRNA dis-regulated in several tumor types and, in colorectal cancer, it plays a critical role in tumor onset, progression, and metastasis. In this paper, we found, that in colorectal cancer cells, the long non-coding RNA H19 can bind immature RNAs and splicing factors as hnRNPM and RBFOX2. Through bioinformatic analysis, we identified 57 transcripts associated with lncH19 and containing binding sites for both splicing factors, hnRNPM, and RBFOX2. Among these transcripts, we identified the mRNA of the GTPase-RAC1, whose alternatively spliced isoform, RAC1B, has been ascribed several roles in the malignant transformation. We confirmed, in vitro, the binding of the splicing factors to both the transcripts RAC1 and lncH19. Loss and gain of expression experiments in two colorectal cancer cell lines (SW620 and HCT116) demonstrated that lncH19 is required for RAC1B expression and, through RAC1B, it induces c-Myc and Cyclin-D increase. In vivo, investigation from biopsies of colorectal cancer patients showed higher levels of all the explored genes (lncH19, RAC1B, c-Myc and Cyclin-D) concerning the healthy counterpart, thus supporting our in vitro model. In addition, we identified a positive correlation between lncH19 and RAC1B in colorectal cancer patients. Finally, we demonstrated that lncH19, as a shuttle, drives the splicing factors RBFOX2 and hnRNPM to RAC1 allowing exon retention and RAC1B expression. The data shown in this paper represent the first evidence of a new mechanism of action by which lncH19 carries out its functions as an oncogene by prompting colorectal cancer through the modulation of alternative splicing.
Keywords: Alternative splicing; Colorectal cancer; RAC1; RBFOX2; RNA-binding proteins; lncH19.
© 2024. The Author(s).
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

