The role of tRNA identity elements in aminoacyl-tRNA editing
- PMID: 39101037
- PMCID: PMC11295145
- DOI: 10.3389/fmicb.2024.1437528
The role of tRNA identity elements in aminoacyl-tRNA editing
Abstract
The rules of the genetic code are implemented by the unique features that define the amino acid identity of each transfer RNA (tRNA). These features, known as "identity elements," mark tRNAs for recognition by aminoacyl-tRNA synthetases (ARSs), the enzymes responsible for ligating amino acids to tRNAs. While tRNA identity elements enable stringent substrate selectivity of ARSs, these enzymes are prone to errors during amino acid selection, leading to the synthesis of incorrect aminoacyl-tRNAs that jeopardize the fidelity of protein synthesis. Many error-prone ARSs have evolved specialized domains that hydrolyze incorrectly synthesized aminoacyl-tRNAs. These domains, known as editing domains, also exist as free-standing enzymes and, together with ARSs, safeguard protein synthesis fidelity. Here, we discuss how the same identity elements that define tRNA aminoacylation play an integral role in aminoacyl-tRNA editing, synergistically ensuring the correct translation of genetic information into proteins. Moreover, we review the distinct strategies of tRNA selection used by editing enzymes and ARSs to avoid undesired hydrolysis of correctly aminoacylated tRNAs.
Keywords: aminoacyl-tRNA synthetases; editing; mistranslation; protein synthesis; tRNA; translational fidelity.
Copyright © 2024 Cruz and Vargas-Rodriguez.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
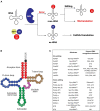

References
Publication types
LinkOut - more resources
Full Text Sources

