Saponin nanoparticle adjuvants incorporating Toll-like receptor agonists drive distinct immune signatures and potent vaccine responses
- PMID: 39110802
- PMCID: PMC11305391
- DOI: 10.1126/sciadv.adn7187
Saponin nanoparticle adjuvants incorporating Toll-like receptor agonists drive distinct immune signatures and potent vaccine responses
Abstract
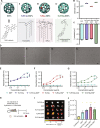
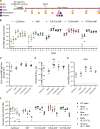
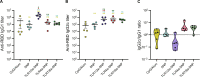
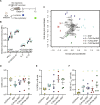
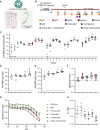
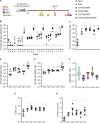
Over the past few decades, the development of potent and safe immune-activating adjuvant technologies has become the heart of intensive research in the constant fight against highly mutative and immune evasive viruses such as influenza, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), and human immunodeficiency virus (HIV). Herein, we developed a highly modular saponin-based nanoparticle platform incorporating Toll-like receptor agonists (TLRas) including TLR1/2a, TLR4a, and TLR7/8a adjuvants and their mixtures. These various TLRa-saponin nanoparticle adjuvant constructs induce unique acute cytokine and immune-signaling profiles, leading to specific T helper responses that could be of interest depending on the target disease for prevention. In a murine vaccine study, the adjuvants greatly improved the potency, durability, breadth, and neutralization of both COVID-19 and HIV vaccine candidates, suggesting the potential broad application of these adjuvant constructs to a range of different antigens. Overall, this work demonstrates a modular TLRa-SNP adjuvant platform that could improve the design of vaccines and affect modern vaccine development.
Figures

Update of
-
Saponin Nanoparticle Adjuvants Incorporating Toll-Like Receptor Agonists Drive Distinct Immune Signatures and Potent Vaccine Responses.bioRxiv [Preprint]. 2024 Jun 27:2023.07.16.549249. doi: 10.1101/2023.07.16.549249. bioRxiv. 2024. Update in: Sci Adv. 2024 Aug 9;10(32):eadn7187. doi: 10.1126/sciadv.adn7187. PMID: 37577608 Free PMC article. Updated. Preprint.
Similar articles
-
Saponin Nanoparticle Adjuvants Incorporating Toll-Like Receptor Agonists Drive Distinct Immune Signatures and Potent Vaccine Responses.bioRxiv [Preprint]. 2024 Jun 27:2023.07.16.549249. doi: 10.1101/2023.07.16.549249. bioRxiv. 2024. Update in: Sci Adv. 2024 Aug 9;10(32):eadn7187. doi: 10.1126/sciadv.adn7187. PMID: 37577608 Free PMC article. Updated. Preprint.
-
TLR7-Adjuvanted Ionizable Lipid Nanoparticles for mRNA Vaccine Delivery.AAPS J. 2025 Apr 25;27(4):80. doi: 10.1208/s12248-025-01073-2. AAPS J. 2025. PMID: 40281311
-
STING agonists as promising vaccine adjuvants to boost immunogenicity against SARS-related coronavirus derived infection: possible role of autophagy.Cell Commun Signal. 2024 Jun 3;22(1):305. doi: 10.1186/s12964-024-01680-0. Cell Commun Signal. 2024. PMID: 38831299 Free PMC article. Review.
-
Design, Synthesis, and Biological Evaluation of New 2,6,7-Substituted Purine Derivatives as Toll-like Receptor 7 Agonists for Intranasal Vaccine Adjuvants.J Med Chem. 2024 Jun 13;67(11):9389-9405. doi: 10.1021/acs.jmedchem.4c00489. Epub 2024 May 24. J Med Chem. 2024. PMID: 38787938
-
Advancing mRNA vaccines: A comprehensive review of design, delivery, and efficacy in infectious diseases.Int J Biol Macromol. 2025 Aug;319(Pt 3):145501. doi: 10.1016/j.ijbiomac.2025.145501. Epub 2025 Jun 24. Int J Biol Macromol. 2025. PMID: 40571008 Review.
Cited by
-
Synergistic Potential of Nanomedicine in Prostate Cancer Immunotherapy: Breakthroughs and Prospects.Int J Nanomedicine. 2024 Oct 2;19:9459-9486. doi: 10.2147/IJN.S466396. eCollection 2024. Int J Nanomedicine. 2024. PMID: 39371481 Free PMC article. Review.
-
Sustained exposure to multivalent antigen-decorated nanoparticles generates broad anti-coronavirus responses.Matter. 2025 Apr 2;8(4):102006. doi: 10.1016/j.matt.2025.102006. Epub 2025 Feb 25. Matter. 2025. PMID: 40821443
-
Recent advances in therapeutic cancer vaccines.Nat Rev Cancer. 2025 Jul;25(7):517-533. doi: 10.1038/s41568-025-00820-z. Epub 2025 May 16. Nat Rev Cancer. 2025. PMID: 40379970 Review.
-
Beyond the Pandemic Era: Recent Advances and Efficacy of SARS-CoV-2 Vaccines Against Emerging Variants of Concern.Vaccines (Basel). 2025 Apr 17;13(4):424. doi: 10.3390/vaccines13040424. Vaccines (Basel). 2025. PMID: 40333293 Free PMC article. Review.
-
Preliminary results on novel adjuvant combinations suggest enhanced immunogenicity of whole inactivated pandemic influenza vaccines.Front Drug Deliv. 2024 Jul 16;4:1382266. doi: 10.3389/fddev.2024.1382266. eCollection 2024. Front Drug Deliv. 2024. PMID: 40836981 Free PMC article.
References
-
- Moyle P. M., Toth I., Modern subunit vaccines: Development, components, and research opportunities. ChemMedChem 8, 360–376 (2013). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

