Fdo1, Fkh1, Fkh2, and the Swi6-Mbp1 MBF complex regulate Mcd1 levels to impact eco1 rad61 cell growth in Saccharomyces cerevisiae
- PMID: 39110836
- PMCID: PMC11457938
- DOI: 10.1093/genetics/iyae128
Fdo1, Fkh1, Fkh2, and the Swi6-Mbp1 MBF complex regulate Mcd1 levels to impact eco1 rad61 cell growth in Saccharomyces cerevisiae
Abstract
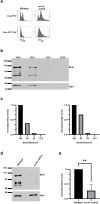
Cohesins promote proper chromosome segregation, gene transcription, genomic architecture, DNA condensation, and DNA damage repair. Mutations in either cohesin subunits or regulatory genes can give rise to severe developmental abnormalities (such as Robert Syndrome and Cornelia de Lange Syndrome) and also are highly correlated with cancer. Despite this, little is known about cohesin regulation. Eco1 (ESCO2/EFO2 in humans) and Rad61 (WAPL in humans) represent two such regulators but perform opposing roles. Eco1 acetylation of cohesin during S phase, for instance, stabilizes cohesin-DNA binding to promote sister chromatid cohesion. On the other hand, Rad61 promotes the dissociation of cohesin from DNA. While Eco1 is essential, ECO1 and RAD61 co-deletion results in yeast cell viability, but only within a limited temperature range. Here, we report that eco1rad61 cell lethality is due to reduced levels of the cohesin subunit Mcd1. Results from a suppressor screen further reveals that FDO1 deletion rescues the temperature-sensitive (ts) growth defects exhibited by eco1rad61 double mutant cells by increasing Mcd1 levels. Regulation of MCD1 expression, however, appears more complex. Elevated expression of MBP1, which encodes a subunit of the MBF transcription complex, also rescues eco1rad61 cell growth defects. Elevated expression of SWI6, however, which encodes the Mbp1-binding partner of MBF, exacerbates eco1rad61 cell growth and also abrogates the Mpb1-dependent rescue. Finally, we identify two additional transcription factors, Fkh1 and Fkh2, that impact MCD1 expression. In combination, these findings provide new insights into the nuanced and multi-faceted transcriptional pathways that impact MCD1 expression.
Keywords: Cornelia de Lange Syndrome (CdLS); ECO1/ESCO2; Fdo1; Fkh1; Fkh2; MBF (Mbp1 and Swi6); Mcd1/Scc1/RAD21; Rad61/WAPL; Roberts Syndrome (RBS); cohesins.
© The Author(s) 2024. Published by Oxford University Press on behalf of The Genetics Society of America.
Conflict of interest statement
Conflicts of interest The author(s) declare no conflict of interest.
Figures
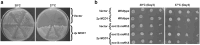



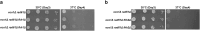


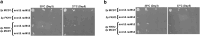



References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

