Genetic modules for α-factor pheromone controlled growth regulation of Saccharomyces cerevisiae
- PMID: 39113811
- PMCID: PMC11300815
- DOI: 10.1002/elsc.202300235
Genetic modules for α-factor pheromone controlled growth regulation of Saccharomyces cerevisiae
Abstract
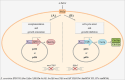
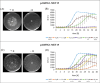
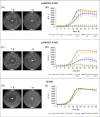
Saccharomyces cerevisiae is a commonly used microorganism in the biotechnological industry. For the industrial heterologous production of compounds, it is of great advantage to work with growth-controllable yeast strains. In our work, we utilized the natural pheromone system of S. cerevisiae and generated a set of different strains possessing an α-pheromone controllable growth behavior. Naturally, the α-factor pheromone is involved in communication between haploid S. cerevisiae cells. Perception of the pheromone initiates several cellular changes, enabling the cells to prepare for an upcoming mating event. We exploited this natural pheromone response system and developed two different plasmid-based modules, in which the target genes, MET15 and FAR1, are under control of the α-factor sensitive FIG1 promoter for a controlled expression in S. cerevisiae. Whereas expression of MET15 led to a growth induction, FAR1 expression inhibited growth. The utilization of low copy number or high copy number plasmids for target gene expression and different concentrations of α-factor allow a finely adjustable control of yeast growth rate.
Keywords: Saccharomyces cerevisiae; growth regulation; pheromone response; synthetic biology; α‐factor.
© 2024 The Author(s). Engineering in Life Sciences published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors have declared no conflicts of interest. The manuscript does not include animal experiments or human studies.
Figures

References
-
- Nielsen J. Yeast systems biology: model organism and cell factory. Biotechnol J. 2019;14:e1800421. - PubMed
-
- Morata A, Loira I, Yeast Industrial Applications, InTech; 2017.
-
- Machida K, Tanaka T, Yano Y, Otani S, Taniguchi M. Farnesol‐induced growth inhibition in Saccharomyces cerevisiae by a cell cycle mechanism. Microbiology. 1999;145:293‐299. - PubMed
LinkOut - more resources
Full Text Sources
