Transcriptomic classification of diffuse large B-cell lymphoma identifies a high-risk activated B-cell-like subpopulation with targetable MYC dysregulation
- PMID: 39117654
- PMCID: PMC11310352
- DOI: 10.1038/s41467-024-50830-y
Transcriptomic classification of diffuse large B-cell lymphoma identifies a high-risk activated B-cell-like subpopulation with targetable MYC dysregulation
Abstract
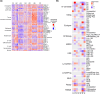
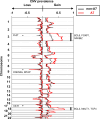
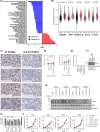
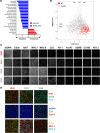
Immunochemotherapy has been the mainstay of treatment for newly diagnosed diffuse large B-cell lymphoma (ndDLBCL) yet is inadequate for many patients. In this work, we perform unsupervised clustering on transcriptomic features from a large cohort of ndDLBCL patients and identify seven clusters, one called A7 with poor prognosis, and develop a classifier to identify these clusters in independent ndDLBCL cohorts. This high-risk cluster is enriched for activated B-cell cell-of-origin, low immune infiltration, high MYC expression, and copy number aberrations. We compare and contrast our methodology with recent DLBCL classifiers to contextualize our clusters and show improved prognostic utility. Finally, using pre-clinical models, we demonstrate a mechanistic rationale for IKZF1/3 degraders such as lenalidomide to overcome the low immune infiltration phenotype of A7 by inducing T-cell trafficking into tumors and upregulating MHC I and II on tumor cells, and demonstrate that TCF4 is an important regulator of MYC-related biology in A7.
© 2024. The Author(s).
Conflict of interest statement
M.E.S., K.W., C.C.H., A.P., M.O., N.S., Y.N., C.H., L.W., H.C., A.S., P.H., and A.K.G. are employees and/or stakeholders in BMS. The other authors declare no competing interests.
Figures



References
MeSH terms
Substances
Grants and funding
- U01 CA195568/CA/NCI NIH HHS/United States
- P30 CA086862/CA/NCI NIH HHS/United States
- CA195568/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
- R03 CA099524/CA/NCI NIH HHS/United States
- CA97274/U.S. Department of Health & Human Services | NIH | National Cancer Institute (NCI)
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

