Spindle architecture constrains karyotype evolution
- PMID: 39117795
- PMCID: PMC11392806
- DOI: 10.1038/s41556-024-01485-w
Spindle architecture constrains karyotype evolution
Abstract
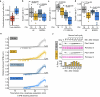
The eukaryotic cell division machinery must rapidly and reproducibly duplicate and partition the cell's chromosomes in a carefully coordinated process. However, chromosome numbers vary dramatically between genomes, even on short evolutionary timescales. We sought to understand how the mitotic machinery senses and responds to karyotypic changes by using a series of budding yeast strains in which the native chromosomes have been successively fused. Using a combination of cell biological profiling, genetic engineering and experimental evolution, we show that chromosome fusions are well tolerated up until a critical point. Cells with fewer than five centromeres lack the necessary number of kinetochore-microtubule attachments needed to counter outward forces in the metaphase spindle, triggering the spindle assembly checkpoint and prolonging metaphase. Our findings demonstrate that spindle architecture is a constraining factor for karyotype evolution.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures






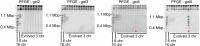


Update of
-
Spindle architecture constrains karyotype in budding yeast.bioRxiv [Preprint]. 2024 May 28:2023.10.25.563899. doi: 10.1101/2023.10.25.563899. bioRxiv. 2024. Update in: Nat Cell Biol. 2024 Sep;26(9):1496-1503. doi: 10.1038/s41556-024-01485-w. PMID: 37961714 Free PMC article. Updated. Preprint.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

