Proteomic aging clock predicts mortality and risk of common age-related diseases in diverse populations
- PMID: 39117878
- PMCID: PMC11405266
- DOI: 10.1038/s41591-024-03164-7
Proteomic aging clock predicts mortality and risk of common age-related diseases in diverse populations
Abstract
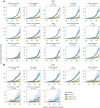
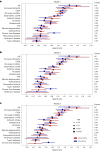
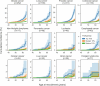
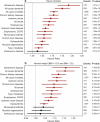
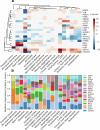
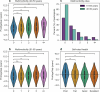
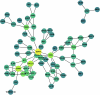
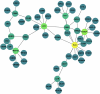
Circulating plasma proteins play key roles in human health and can potentially be used to measure biological age, allowing risk prediction for age-related diseases, multimorbidity and mortality. Here we developed a proteomic age clock in the UK Biobank (n = 45,441) using a proteomic platform comprising 2,897 plasma proteins and explored its utility to predict major disease morbidity and mortality in diverse populations. We identified 204 proteins that accurately predict chronological age (Pearson r = 0.94) and found that proteomic aging was associated with the incidence of 18 major chronic diseases (including diseases of the heart, liver, kidney and lung, diabetes, neurodegeneration and cancer), as well as with multimorbidity and all-cause mortality risk. Proteomic aging was also associated with age-related measures of biological, physical and cognitive function, including telomere length, frailty index and reaction time. Proteins contributing most substantially to the proteomic age clock are involved in numerous biological functions, including extracellular matrix interactions, immune response and inflammation, hormone regulation and reproduction, neuronal structure and function and development and differentiation. In a validation study involving biobanks in China (n = 3,977) and Finland (n = 1,990), the proteomic age clock showed similar age prediction accuracy (Pearson r = 0.92 and r = 0.94, respectively) compared to its performance in the UK Biobank. Our results demonstrate that proteomic aging involves proteins spanning multiple functional categories and can be used to predict age-related functional status, multimorbidity and mortality risk across geographically and genetically diverse populations.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures






References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

