Identification of Ribonuclease Inhibitors for the Control of Pathogenic Bacteria
- PMID: 39125622
- PMCID: PMC11311990
- DOI: 10.3390/ijms25158048
Identification of Ribonuclease Inhibitors for the Control of Pathogenic Bacteria
Abstract
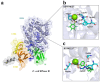
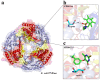
Bacteria are known to be constantly adapting to become resistant to antibiotics. Currently, efficient antibacterial compounds are still available; however, it is only a matter of time until these compounds also become inefficient. Ribonucleases are the enzymes responsible for the maturation and degradation of RNA molecules, and many of them are essential for microbial survival. Members of the PNPase and RNase II families of exoribonucleases have been implicated in virulence in many pathogens and, as such, are valid targets for the development of new antibacterials. In this paper, we describe the use of virtual high-throughput screening (vHTS) to identify chemical compounds predicted to bind to the active sites within the known structures of RNase II and PNPase from Escherichia coli. The subsequent in vitro screening identified compounds that inhibited the activity of these exoribonucleases, with some also affecting cell viability, thereby providing proof of principle for utilizing the known structures of these enzymes in the pursuit of new antibacterials.
Keywords: PNPase; RNase II; RNase R; antimicrobials; virtual high-throughput screening (vHTS).
Conflict of interest statement
The authors declare no conflicts of interest.
Figures




Similar articles
-
The first small-molecule inhibitors of members of the ribonuclease E family.Sci Rep. 2015 Jan 26;5:8028. doi: 10.1038/srep08028. Sci Rep. 2015. PMID: 25619596 Free PMC article.
-
Next generation sequencing analysis reveals that the ribonucleases RNase II, RNase R and PNPase affect bacterial motility and biofilm formation in E. coli.BMC Genomics. 2015 Feb 14;16(1):72. doi: 10.1186/s12864-015-1237-6. BMC Genomics. 2015. PMID: 25757888 Free PMC article.
-
RNase activity of polynucleotide phosphorylase is critical at low temperature in Escherichia coli and is complemented by RNase II.J Bacteriol. 2008 Sep;190(17):5924-33. doi: 10.1128/JB.00500-08. Epub 2008 Jul 7. J Bacteriol. 2008. PMID: 18606734 Free PMC article.
-
Major 3'-5' Exoribonucleases in the Metabolism of Coding and Non-coding RNA.Prog Mol Biol Transl Sci. 2018;159:101-155. doi: 10.1016/bs.pmbts.2018.07.005. Epub 2018 Aug 25. Prog Mol Biol Transl Sci. 2018. PMID: 30340785 Review.
-
The role of 3'-5' exoribonucleases in RNA degradation.Prog Mol Biol Transl Sci. 2009;85:187-229. doi: 10.1016/S0079-6603(08)00805-2. Prog Mol Biol Transl Sci. 2009. PMID: 19215773 Review.
Cited by
-
Carrot and stick: how RNase R contributes to function and destruction of the translation machinery.RNA Biol. 2025 Dec;22(1):1-22. doi: 10.1080/15476286.2025.2535846. Epub 2025 Jul 29. RNA Biol. 2025. PMID: 40734258 Free PMC article. Review.
References
-
- Mardle C.E., Goddard L.R., Spelman B.C., Atkins H.S., Butt L.E., Cox P.A., Gowers D.M., Vincent H.A., Callaghan A.J. Identification and analysis of novel small molecule inhibitors of RNase E: Implications for antibacterial targeting and regulation of RNase E. Biochem. Biophys. Rep. 2020;23:100773. doi: 10.1016/j.bbrep.2020.100773. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

