Immune landscape of isocitrate dehydrogenase-stratified primary and recurrent human gliomas
- PMID: 39126294
- PMCID: PMC11630528
- DOI: 10.1093/neuonc/noae139
Immune landscape of isocitrate dehydrogenase-stratified primary and recurrent human gliomas
Abstract
Background: Human gliomas are classified using isocitrate dehydrogenase (IDH) status as a prognosticator; however, the influence of genetic differences and treatment effects on ensuing immunity remains unclear.
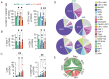
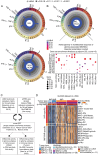
Methods: In this study, we used sequential single-cell transcriptomics on 144 678 and spectral cytometry on over 2 million immune cells encompassing 48 human gliomas to decipher their immune landscape.
Results: We identified 22 distinct immune cell types that contribute to glioma immunity. Specifically, brain-resident microglia (MG) were reduced with a concomitant increase in CD8+ T lymphocytes during glioma recurrence independent of IDH status. In contrast, IDH-wild type-associated patterns, such as an abundance of antigen-presenting cell-like MG and cytotoxic CD8+ T cells, were observed. Beyond elucidating the differences in IDH, relapse, and treatment-associated immunity, we discovered novel inflammatory MG subpopulations expressing granulysin, a cytotoxic peptide that is otherwise expressed in lymphocytes only. Furthermore, we provide a robust genomic framework for defining macrophage polarization beyond M1/M2 paradigm and reference signatures of glioma-specific tumor immune microenvironment (termed GlioTIME-36) for deconvoluting transcriptomic datasets.
Conclusions: This study provides advanced optics of the human pan-glioma immune contexture as a valuable guide for translational and clinical applications.
Keywords: glioma; isocitrate dehydrogenase; microglia; tumor immune microenvironment.
© The Author(s) 2024. Published by Oxford University Press on behalf of the Society for Neuro-Oncology. All rights reserved. For commercial re-use, please contact reprints@oup.com for reprints and translation rights for reprints. All other permissions can be obtained through our RightsLink service via the Permissions link on the article page on our site—for further information please contact journals.permissions@oup.com.
Conflict of interest statement
All authors declare no competing interests except H.S., S.A.S., E.R.P.C., and A.B.H. H.S. is a founder of NextGen Omics. S.A.S is engaged in consulting with Boston Scientific, Neuropace, Zimmer Biomet, Koh Young, Sensoria Therapeutics, and Varian Medical. E.R.P.C. executed consulting assignments for Nuclei Lt., iTeos Belgium. A.B.H. has engaged in contracts with Abbvie, Ainylam, Codiak, Cellularity and received royalties from DNAtrix, Celldex Therapeutics and consulting fees from Novocure, Istari Oncology, Alphasights, and BlueRock Therapeutics. A.B.H. is an advisory board member with WCG Oncology, Caris Life Science, Children’s National Hospital Brain Tumor Institute, UCSF Neurological and Brain Tumor Program, Dana Farber and Brigham and Women’s Hospital (P01), Cleveland Clinic Sex Difference (P01), UCLA Brain SPORE, National Cancer Advisory Board. She has stocks in Caris Life Sciences and is a recipient of gifts and other services from Moleculin, Carthera, and Takeda.
Figures






References
-
- Leng F, Edison P.. Neuroinflammation and microglial activation in Alzheimer disease: where do we go from here? Nat Rev Neurol. 2021;17(3):157–172. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

