This is a preprint.
Loss of RREB1 reduces adipogenesis and improves insulin sensitivity in mouse and human adipocytes
- PMID: 39131393
- PMCID: PMC11312556
- DOI: 10.1101/2024.07.30.605923
Loss of RREB1 reduces adipogenesis and improves insulin sensitivity in mouse and human adipocytes
Abstract
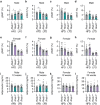
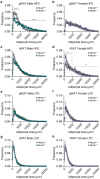
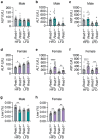
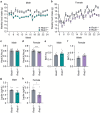
There are multiple independent genetic signals at the Ras-responsive element binding protein 1 (RREB1) locus associated with type 2 diabetes risk, fasting glucose, ectopic fat, height, and bone mineral density. We have previously shown that loss of RREB1 in pancreatic beta cells reduces insulin content and impairs islet cell development and function. However, RREB1 is a widely expressed transcription factor and the metabolic impact of RREB1 loss in vivo remains unknown. Here, we show that male and female global heterozygous knockout (Rreb1 +/-) mice have reduced body length, weight, and fat mass on high-fat diet. Rreb1+/- mice have sex- and diet-specific decreases in adipose tissue and adipocyte size; male mice on high-fat diet had larger gonadal adipocytes, while males on standard chow and females on high-fat diet had smaller, more insulin sensitive subcutaneous adipocytes. Mouse and human precursor cells lacking RREB1 have decreased adipogenic gene expression and activated transcription of genes associated with osteoblast differentiation, which was associated with Rreb1 +/- mice having increased bone mineral density in vivo. Finally, human carriers of RREB1 T2D protective alleles have smaller adipocytes, consistent with RREB1 loss-of-function reducing diabetes risk.
Keywords: Diabetes; RREB1; adipocyte; insulin sensitivity; transcription factor.
Conflict of interest statement
Competing interests ALG discloses that her spouse is an employee of Genentech and hold stock options in Roche. All other authors declare no interests that could be considered conflicting.
Figures







References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
