A computational approach to developing a multi-epitope vaccine for combating Pseudomonas aeruginosa-induced pneumonia and sepsis
- PMID: 39133098
- PMCID: PMC11318047
- DOI: 10.1093/bib/bbae401
A computational approach to developing a multi-epitope vaccine for combating Pseudomonas aeruginosa-induced pneumonia and sepsis
Abstract
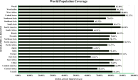
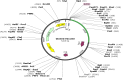
Pseudomonas aeruginosa is a complex nosocomial infectious agent responsible for numerous illnesses, with its growing resistance variations complicating treatment development. Studies have emphasized the importance of virulence factors OprE and OprF in pathogenesis, highlighting their potential as vaccine candidates. In this study, B-cell, MHC-I, and MHC-II epitopes were identified, and molecular linkers were active to join these epitopes with an appropriate adjuvant to construct a vaccine. Computational tools were employed to forecast the tertiary framework, characteristics, and also to confirm the vaccine's composition. The potency was weighed through population coverage analysis and immune simulation. This project aims to create a multi-epitope vaccine to reduce P. aeruginosa-related illness and mortality using immunoinformatics resources. The ultimate complex has been determined to be stable, soluble, antigenic, and non-allergenic upon inspection of its physicochemical and immunological properties. Additionally, the protein exhibited acidic and hydrophilic characteristics. The Ramachandran plot, ProSA-web, ERRAT, and Verify3D were employed to ensure the final model's authenticity once the protein's three-dimensional structure had been established and refined. The vaccine model showed a significant binding score and stability when interacting with MHC receptors. Population coverage analysis indicated a global coverage rate of 83.40%, with the USA having the highest coverage rate, exceeding 90%. Moreover, the vaccine sequence underwent codon optimization before being cloned into the Escherichia coli plasmid vector pET-28a (+) at the EcoRI and EcoRV restriction sites. Our research has developed a vaccine against P. aeruginosa that has strong binding affinity and worldwide coverage, offering an acceptable way to mitigate nosocomial infections.
Keywords: OprE and OprF; Pseudomonas aeruginosa; immunoinformatics; multi-epitope vaccine.
© The Author(s) 2024. Published by Oxford University Press.
Figures




Similar articles
-
Designing a multi-epitope vaccine against Pseudomonas aeruginosa via integrating reverse vaccinology with immunoinformatics approaches.Sci Rep. 2025 Mar 26;15(1):10425. doi: 10.1038/s41598-025-90226-6. Sci Rep. 2025. PMID: 40140433 Free PMC article.
-
Whole proteome-integrated and vaccinomics-based next generation mRNA vaccine design against Pseudomonas aeruginosa-A hierarchical subtractive proteomics approach.Int J Biol Macromol. 2025 May;309(Pt 1):142627. doi: 10.1016/j.ijbiomac.2025.142627. Epub 2025 Mar 31. Int J Biol Macromol. 2025. PMID: 40174835
-
Immunoinformatics-aided design of a new multi-epitope vaccine adjuvanted with domain 4 of pneumolysin against Streptococcus pneumoniae strains.BMC Bioinformatics. 2023 Feb 24;24(1):67. doi: 10.1186/s12859-023-05175-6. BMC Bioinformatics. 2023. PMID: 36829109 Free PMC article.
-
Pseudomonas aeruginosa antigens as potential vaccines.FEMS Microbiol Rev. 1997 Nov;21(3):243-77. doi: 10.1111/j.1574-6976.1997.tb00353.x. FEMS Microbiol Rev. 1997. PMID: 9451816 Review.
-
Potential of protein OprF of Pseudomonas in bivalent vaccines.Behring Inst Mitt. 1997 Feb;(98):283-90. Behring Inst Mitt. 1997. PMID: 9382752 Review.
Cited by
-
Designing a broad-spectrum multi-epitope subunit vaccine against leptospirosis using immunoinformatics and structural approaches.Front Immunol. 2025 Jan 28;15:1503853. doi: 10.3389/fimmu.2024.1503853. eCollection 2024. Front Immunol. 2025. PMID: 39936152 Free PMC article.
-
Designing a multi-epitope vaccine against Pseudomonas aeruginosa via integrating reverse vaccinology with immunoinformatics approaches.Sci Rep. 2025 Mar 26;15(1):10425. doi: 10.1038/s41598-025-90226-6. Sci Rep. 2025. PMID: 40140433 Free PMC article.
-
Towards precision epitopes based vaccine against Enterococcus faecalis by integrating vaccinomics, reverse vaccinology and biophysics approaches.Biochem Biophys Rep. 2025 Jun 10;43:102082. doi: 10.1016/j.bbrep.2025.102082. eCollection 2025 Sep. Biochem Biophys Rep. 2025. PMID: 40546343 Free PMC article.
-
Peptide vaccines: an innovative therapeutic approach against antibiotic-resistant bacterial infections.Front Immunol. 2025 May 21;16:1567584. doi: 10.3389/fimmu.2025.1567584. eCollection 2025. Front Immunol. 2025. PMID: 40469313 Free PMC article. Review.
References
-
- Tacconelli E, Carrara E, Savoldi A. et al. . Articles discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. 2017. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

