Complete mitochondrial genome of Acanthochitona defilippii (Polyplacophora: Chitonida) from South Korea
- PMID: 39135642
- PMCID: PMC11318481
- DOI: 10.1080/23802359.2024.2386403
Complete mitochondrial genome of Acanthochitona defilippii (Polyplacophora: Chitonida) from South Korea
Abstract
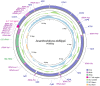
The chiton (Polyplacophora) occupies a significant position in molluscan evolutionary history as one of the most primitive groups within the phylum Mollusca. Acanthochitona defilippii (Tapparone-Canefri 1874) (Chitonida: Acanthochitonidae) is a commonly found intertidal chiton species in South Korea. In this study, we characterized the complete mitochondrial genome of A. defilippii (14,999 bp long), comprising 13 protein-coding genes (PCGs), 22 transfer RNA genes, two ribosomal RNA genes, and an A + T rich region (166 bp). The base composition is as follows: 31.82% for A, 11.63% for C, 16.69% for G, and 39.86% for T. We reconstructed a maximum likelihood (ML) tree to elucidate phylogenetic relationships among the eight chitonid families using the nucleotide sequences of all PCGs. The ML tree revealed that A. defilippii clustered with Acanthochitona avicula (BP 100) within the family Acanthochitonidae. Acanthochitonidae formed a sister group with Mopaliidae. The results could provide a valuable understanding the phylogenetic relationships of chitonid species.
Keywords: Acanthochitona defilippii; Acanthochitonidae; chiton; mitochondrial genome; phylogeny.
© 2024 The Author(s). Published by Informa UK Limited, trading as Taylor & Francis Group.
Conflict of interest statement
The authors report no conflicts of interest. The authors alone are responsible for the content and writing of the paper.
Figures


References
-
- Alnashiri H, Thomas L, Philip S, Thaikkottathil M, Sureshkumar S, Kutty R.. 2024. Complete mitochondrial genome and phylogenetic relationships of the red sea Chiton Acanthopleura vaillantii Rochebrune, 1882 (Polyplacophora: chitonida). Thalassas. 40(1):51–58. doi:10.1007/s41208-023-00648-0. - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources
