HLTF resolves G4s and promotes G4-induced replication fork slowing to maintain genome stability
- PMID: 39142279
- PMCID: PMC11366124
- DOI: 10.1016/j.molcel.2024.07.018
HLTF resolves G4s and promotes G4-induced replication fork slowing to maintain genome stability
Abstract
G-quadruplexes (G4s) form throughout the genome and influence important cellular processes. Their deregulation can challenge DNA replication fork progression and threaten genome stability. Here, we demonstrate an unexpected role for the double-stranded DNA (dsDNA) translocase helicase-like transcription factor (HLTF) in responding to G4s. We show that HLTF, which is enriched at G4s in the human genome, can directly unfold G4s in vitro and uses this ATP-dependent translocase function to suppress G4 accumulation throughout the cell cycle. Additionally, MSH2 (a component of MutS heterodimers that bind G4s) and HLTF act synergistically to suppress G4 accumulation, restrict alternative lengthening of telomeres, and promote resistance to G4-stabilizing drugs. In a discrete but complementary role, HLTF restrains DNA synthesis when G4s are stabilized by suppressing primase-polymerase (PrimPol)-dependent repriming. Together, the distinct roles of HLTF in the G4 response prevent DNA damage and potentially mutagenic replication to safeguard genome stability.
Keywords: DNA replication stress response; DNA translocase; G-quadruplex; HLTF; MSH2; PrimPol; RNA-DNA hybrid; alternative lengthening of telomeres; genome stability; nucleic acid secondary structure.
Copyright © 2024 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests K.A.C. is a member of the scientific advisory board of IDEAYA Biosciences and RADD Pharmaceuticals and is on the oncology advisory board for GlaxoSmithKline. S.J.B. is a co-founder, VP Science Strategy and shareholder at Artios Pharma Ltd. K.A.C. and S.J.B. are also members of the advisory board at Molecular Cell.
Figures



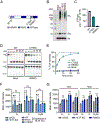



Update of
-
HLTF Prevents G4 Accumulation and Promotes G4-induced Fork Slowing to Maintain Genome Stability.bioRxiv [Preprint]. 2023 Oct 27:2023.10.27.563641. doi: 10.1101/2023.10.27.563641. bioRxiv. 2023. Update in: Mol Cell. 2024 Aug 22;84(16):3044-3060.e11. doi: 10.1016/j.molcel.2024.07.018. PMID: 37961428 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

