This is a preprint.
Adenomas from individuals with pathogenic biallelic variants in the MUTYH and NTHL1 genes demonstrate base excision repair tumour mutational signature profiles similar to colorectal cancers, expanding potential diagnostic and variant classification applications
- PMID: 39148833
- PMCID: PMC11326331
- DOI: 10.1101/2024.08.08.24311713
Adenomas from individuals with pathogenic biallelic variants in the MUTYH and NTHL1 genes demonstrate base excision repair tumour mutational signature profiles similar to colorectal cancers, expanding potential diagnostic and variant classification applications
Update in
-
Adenomas from individuals with pathogenic biallelic variants in the MUTYH and NTHL1 genes demonstrate base excision repair tumour mutational signature profiles similar to colorectal cancers, expanding potential diagnostic and variant classification applications.Transl Oncol. 2025 Feb;52:102266. doi: 10.1016/j.tranon.2024.102266. Epub 2025 Jan 9. Transl Oncol. 2025. PMID: 39793275 Free PMC article.
Abstract
Background: Colorectal cancers (CRCs) from people with biallelic germline likely pathogenic/pathogenic variants in MUTYH or NTHL1 exhibit specific single base substitution (SBS) mutational signatures, namely combined SBS18 and SBS36 (SBS18+SBS36), and SBS30, respectively. The aim was to determine if adenomas from biallelic cases demonstrated these mutational signatures at diagnostic levels.
Methods: Whole-exome sequencing of FFPE tissue and matched blood-derived DNA was performed on 9 adenomas and 15 CRCs from 13 biallelic MUTYH cases, on 7 adenomas and 2 CRCs from 5 biallelic NTHL1 cases and on 27 adenomas and 26 CRCs from 46 non-hereditary (sporadic) participants. All samples were assessed for COSMIC v3.2 SBS mutational signatures.
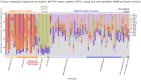
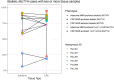
Results: In biallelic MUTYH cases, SBS18+SBS36 signature proportions in adenomas (mean±standard deviation, 65.6%±29.6%) were not significantly different to those observed in CRCs (76.2%±20.5%, p-value=0.37), but were significantly higher compared with non-hereditary adenomas (7.6%±7.0%, p-value=3.4×10-4). Similarly, in biallelic NTHL1 cases, SBS30 signature proportions in adenomas (74.5%±9.4%) were similar to those in CRCs (78.8%±2.4%) but significantly higher compared with non-hereditary adenomas (2.8%±3.6%, p-value=5.1×10-7). Additionally, a compound heterozygote with the c.1187G>A p.(Gly396Asp) pathogenic variant and the c.533G>C p.(Gly178Ala) variant of unknown significance (VUS) in MUTYH demonstrated high levels of SBS18+SBS36 in four adenomas and one CRC, providing evidence for reclassification of the VUS to pathogenic.
Conclusions: SBS18+SBS36 and SBS30 were enriched in adenomas at comparable proportions observed in CRCs from biallelic MUTYH and biallelic NTHL1 cases, respectively. Therefore, testing adenomas may improve the identification of biallelic cases and facilitate variant classification, ultimately enabling opportunities for CRC prevention.
Keywords: Colorectal cancer; MUTYH; NTHL1; SBS18; SBS30; SBS36; adenoma; hereditary cancer predisposition; mutational signature; variant of uncertain clinical significance.
Conflict of interest statement
Robert C. Grant received a scholarship from Pfizer and provided consulting or advisory roles for Astrazeneca, Tempus, Eisai, Incyte, Knight Therapeutics, Guardant Health, and Ipsen. All other authors have no relevant financial or non-financial interests to disclose.
Figures

References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
