Kick-starting the zygotic genome: licensors, specifiers, and beyond
- PMID: 39160344
- PMCID: PMC11467316
- DOI: 10.1038/s44319-024-00223-5
Kick-starting the zygotic genome: licensors, specifiers, and beyond
Abstract
Zygotic genome activation (ZGA), the first transcription event following fertilization, kickstarts the embryonic program that takes over the control of early development from the maternal products. How ZGA occurs, especially in mammals, is poorly understood due to the limited amount of research materials. With the rapid development of single-cell and low-input technologies, remarkable progress made in the past decade has unveiled dramatic transitions of the epigenomes, transcriptomes, proteomes, and metabolomes associated with ZGA. Moreover, functional investigations are yielding insights into the key regulators of ZGA, among which two major classes of players are emerging: licensors and specifiers. Licensors would control the permission of transcription and its timing during ZGA. Accumulating evidence suggests that such licensors of ZGA include regulators of the transcription apparatus and nuclear gatekeepers. Specifiers would instruct the activation of specific genes during ZGA. These specifiers include key transcription factors present at this stage, often facilitated by epigenetic regulators. Based on data primarily from mammals but also results from other species, we discuss in this review how recent research sheds light on the molecular regulation of ZGA and its executors, including the licensors and specifiers.
Keywords: Early Embryo; Licensor; Specifier; Transcription Factor; ZGA.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



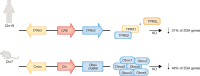
References
-
- Abe K, Schauer T, Torres-Padilla ME (2022) Distinct patterns of RNA polymerase II and transcriptional elongation characterize mammalian genome activation. Cell Rep 41:111865 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 31988101/MOST | National Natural Science Foundation of China (NSFC)
- 31830047/MOST | National Natural Science Foundation of China (NSFC)
- 31725018/MOST | National Natural Science Foundation of China (NSFC)
- 2021YFA1100102/MOST | National Key Research and Development Program of China (NKPs)
- 2019YFA0508900/MOST | National Key Research and Development Program of China (NKPs)
LinkOut - more resources
Full Text Sources
Miscellaneous

