CsPHRs-CsJAZ3 incorporates phosphate signaling and jasmonate pathway to regulate catechin biosynthesis in Camellia sinensis
- PMID: 39161738
- PMCID: PMC11331543
- DOI: 10.1093/hr/uhae178
CsPHRs-CsJAZ3 incorporates phosphate signaling and jasmonate pathway to regulate catechin biosynthesis in Camellia sinensis
Abstract
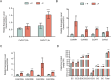
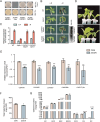
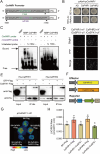
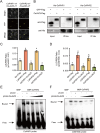
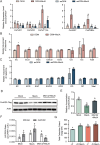
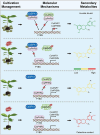
Catechins constitute abundant metabolites in tea and have potential health benefits and high economic value. Intensive study has shown that the biosynthesis of tea catechins is regulated by environmental factors and hormonal signals. However, little is known about the coordination of phosphate (Pi) signaling and the jasmonic acid (JA) pathway on biosynthesis of tea catechins. We found that Pi deficiency caused changes in the content of catechins and modulated the expression levels of genes involved in catechin biosynthesis. Herein, we identified two transcription factors of phosphate signaling in tea, named CsPHR1 and CsPHR2, respectively. Both regulated catechin biosynthesis by activating the transcription of CsANR1 and CsMYB5c. We further demonstrated CsSPX1, a Pi pathway repressor, suppressing the activation by CsPHR1/2 of CsANR1 and CsMYB5c. JA, one of the endogenous plant hormones, has been reported to be involved in the regulation of secondary metabolism. Our work demonstrated that the JA signaling repressor CsJAZ3 negatively regulated catechin biosynthesis via physical interaction with CsPHR1 and CsPHR2. Thus, the CsPHRs-CsJAZ3 module bridges the nutrition and hormone signals, contributing to targeted cultivation of high-quality tea cultivars with high fertilizer efficiency.
© The Author(s) 2024. Published by Oxford University Press on behalf of Nanjing Agricultural University.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
References
-
- Fraga CG, Croft KD, Kennedy DO. et al. The effects of polyphenols and other bioactives on human health. Food Funct. 2019;10:514–28 - PubMed
-
- Goya L, Roman RS, de Pascual-Teresa S. Polyphenols' effect on cerebrovascular health. Curr Med Chem. 2022;29:1029–44 - PubMed
-
- Veeraraghavan VP, Mony U, Renu K. et al. Effects of polyphenols on ncRNAs in cancer – an update. Clin Exp Pharmacol Physiol. 2022;49:613–23 - PubMed
-
- Roh E, Kim JE, Kwon JY. et al. Molecular mechanisms of green tea polyphenols with protective effects against skin photoaging. Crit Rev Food Sci Nutr. 2017;57:1631–7 - PubMed
-
- Zhang X, Liu Y, Gao K. et al. Characterisation of anthocyanidin reductase from Shuchazao green tea. J Sci Food Agric. 2012;92:1533–9 - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

