RNA-directed peptide synthesis across a nicked loop
- PMID: 39164017
- PMCID: PMC11514466
- DOI: 10.1093/nar/gkae702
RNA-directed peptide synthesis across a nicked loop
Abstract
Ribosomal translation at the origin of life requires controlled aminoacylation to produce mono-aminoacyl esters of tRNAs. Herein, we show that transient annealing of short RNA oligo:amino acid mixed anhydrides to an acceptor strand enables the sequential transfer of aminoacyl residues to the diol of an overhang, first forming aminoacyl esters then peptidyl esters. Using N-protected aminoacyl esters prevents unwanted peptidyl ester formation in this manner. However, N-acyl-aminoacyl transfer is not stereoselective.
Plain language summary
The Central Dogma dominates our understanding of modern biology. However, the mechanisms of how RNA directed peptide synthesis (translation) developed at the dawn of life remain a puzzle. In this study, we demonstrate that short peptides can spontaneously form at the end of a duplex RNA with an overhang. Contemporary proteins are composed exclusively of L-amino acids and our research reveals that L-amino acids with free amino groups are also more likely to participate in this prebiotic peptide synthesis mechanism. Conversely, when one of the amino acids is protected as it is in modern bacteria, this preference disappears. We demonstrate this chemistry can afford a variety of peptide sequences in a sequence-dependent manner.
© The Author(s) 2024. Published by Oxford University Press on behalf of Nucleic Acids Research.
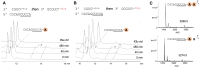
Figures




Similar articles
-
Thioester-mediated RNA aminoacylation and peptidyl-RNA synthesis in water.Nature. 2025 Aug;644(8078):933-944. doi: 10.1038/s41586-025-09388-y. Epub 2025 Aug 27. Nature. 2025. PMID: 40866676 Free PMC article.
-
Interstrand Aminoacyl Transfer in a tRNA Acceptor Stem-Overhang Mimic.J Am Chem Soc. 2021 Aug 4;143(30):11836-11842. doi: 10.1021/jacs.1c05746. Epub 2021 Jul 20. J Am Chem Soc. 2021. PMID: 34283595 Free PMC article.
-
A potential role for RNA aminoacylation prior to its role in peptide synthesis.Proc Natl Acad Sci U S A. 2024 Aug 27;121(35):e2410206121. doi: 10.1073/pnas.2410206121. Epub 2024 Aug 23. Proc Natl Acad Sci U S A. 2024. PMID: 39178230 Free PMC article.
-
Molecular basis for chiral selection in RNA aminoacylation.Int J Mol Sci. 2011;12(7):4745-57. doi: 10.3390/ijms12074745. Epub 2011 Jul 22. Int J Mol Sci. 2011. PMID: 21845109 Free PMC article. Review.
-
The synthesis of peptides and proteins containing non-natural amino acids.Chem Soc Rev. 2004 Sep 10;33(7):422-30. doi: 10.1039/b312953p. Epub 2004 Aug 13. Chem Soc Rev. 2004. PMID: 15354223 Review.
Cited by
-
Purine Chemistry in the Early RNA World at the Origins of Life: From RNA and Nucleobases Lesions to Current Key Metabolic Routes.Chembiochem. 2025 Jun 3;26(11):e202500035. doi: 10.1002/cbic.202500035. Epub 2025 Apr 16. Chembiochem. 2025. PMID: 40237374 Free PMC article. Review.
-
Permeability selection of biologically relevant membranes matches the stereochemistry of life on Earth.PLoS Biol. 2025 May 20;23(5):e3003155. doi: 10.1371/journal.pbio.3003155. eCollection 2025 May. PLoS Biol. 2025. PMID: 40392769 Free PMC article.
References
-
- Krebs J.E., Goldstein E.S., Kilpatrick S.T.. Lewin's Genes XII. 2017; MA: Jones and Bartlett Publishers; 583–620.
-
- Nissen P., Hansen J., Ban N., Moore P.B., Steitz T.A.. The structural basis of ribosome activity in peptide bond synthesis. Science. 2000; 289:920–930. - PubMed
-
- Gilbert W. The origin of life: the RNA world. Nature. 1986; 319:618.
-
- Zhang B., Cech T.R.. Peptide bond formation by in vitro selected ribozymes. Nature. 1997; 390:96–100. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

