Liver eQTL meta-analysis illuminates potential molecular mechanisms of cardiometabolic traits
- PMID: 39173627
- PMCID: PMC11393674
- DOI: 10.1016/j.ajhg.2024.07.017
Liver eQTL meta-analysis illuminates potential molecular mechanisms of cardiometabolic traits
Abstract
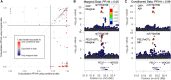
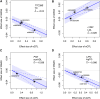
Understanding the molecular mechanisms of complex traits is essential for developing targeted interventions. We analyzed liver expression quantitative-trait locus (eQTL) meta-analysis data on 1,183 participants to identify conditionally distinct signals. We found 9,013 eQTL signals for 6,564 genes; 23% of eGenes had two signals, and 6% had three or more signals. We then integrated the eQTL results with data from 29 cardiometabolic genome-wide association study (GWAS) traits and identified 1,582 GWAS-eQTL colocalizations for 747 eGenes. Non-primary eQTL signals accounted for 17% of all colocalizations. Isolating signals by conditional analysis prior to coloc resulted in 37% more colocalizations than using marginal eQTL and GWAS data, highlighting the importance of signal isolation. Isolating signals also led to stronger evidence of colocalization: among 343 eQTL-GWAS signal pairs in multi-signal regions, analyses that isolated the signals of interest resulted in higher posterior probability of colocalization for 41% of tests. Leveraging allelic heterogeneity, we predicted causal effects of gene expression on liver traits for four genes. To predict functional variants and regulatory elements, we colocalized eQTL with liver chromatin accessibility QTL (caQTL) and found 391 colocalizations, including 73 with non-primary eQTL signals and 60 eQTL signals that colocalized with both a caQTL and a GWAS signal. Finally, we used publicly available massively parallel reporter assays in HepG2 to highlight 14 eQTL signals that include at least one expression-modulating variant. This multi-faceted approach to unraveling the genetic underpinnings of liver-related traits could lead to therapeutic development.
Keywords: GWAS; allelic heterogeneity; colocalization; complex traits; eQTL meta-analysis; liver; signal identification.
Copyright © 2024 American Society of Human Genetics. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests F.I. is an employee of BeiGene and received stocks from the company. F.I. was also an AbbVie employee and received stocks from that company.
Figures



Similar articles
-
Genetic effects on chromatin accessibility uncover mechanisms of liver gene regulation and quantitative traits.Genome Res. 2025 Jul 1;35(7):1485-1502. doi: 10.1101/gr.279741.124. Genome Res. 2025. PMID: 40393811
-
Adipose tissue eQTL meta-analysis highlights the contribution of allelic heterogeneity to gene expression regulation and cardiometabolic traits.Nat Genet. 2025 Jan;57(1):180-192. doi: 10.1038/s41588-024-01982-6. Epub 2025 Jan 2. Nat Genet. 2025. PMID: 39747594 Free PMC article.
-
Multi-omics analysis for identifying cell-type-specific and bulk-level druggable targets in Alzheimer's disease.J Transl Med. 2025 Jul 13;23(1):788. doi: 10.1186/s12967-025-06739-1. J Transl Med. 2025. PMID: 40653482 Free PMC article.
-
Signs and symptoms to determine if a patient presenting in primary care or hospital outpatient settings has COVID-19.Cochrane Database Syst Rev. 2022 May 20;5(5):CD013665. doi: 10.1002/14651858.CD013665.pub3. Cochrane Database Syst Rev. 2022. PMID: 35593186 Free PMC article.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2021 Apr 19;4(4):CD011535. doi: 10.1002/14651858.CD011535.pub4. Cochrane Database Syst Rev. 2021. Update in: Cochrane Database Syst Rev. 2022 May 23;5:CD011535. doi: 10.1002/14651858.CD011535.pub5. PMID: 33871055 Free PMC article. Updated.
Cited by
-
Higher eQTL power reveals signals that boost GWAS colocalization.bioRxiv [Preprint]. 2025 Aug 5:2025.08.05.668745. doi: 10.1101/2025.08.05.668745. bioRxiv. 2025. PMID: 40799583 Free PMC article. Preprint.
-
Accelerating Medicines Partnership in Type 2 Diabetes and Common Metabolic Diseases: Collaborating to Maximize the Value of Genetic and Genomic Data.Diabetes. 2025 Jul 1;74(7):1089-1098. doi: 10.2337/db25-0042. Diabetes. 2025. PMID: 40272257 Free PMC article. Review.
-
Role of expression quantitative trait loci (eQTL) in understanding genetic mechanisms underlying common complex diseases.Mol Cells. 2025 Sep;48(9):100256. doi: 10.1016/j.mocell.2025.100256. Epub 2025 Jul 18. Mol Cells. 2025. PMID: 40684919 Free PMC article. Review.
References
-
- Strunz T., Grassmann F., Gayán J., Nahkuri S., Souza-Costa D., Maugeais C., Fauser S., Nogoceke E., Weber B.H.F. A mega-analysis of expression quantitative trait loci (eQTL) provides insight into the regulatory architecture of gene expression variation in liver. Sci. Rep. 2018;8:5865. - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

