Fc-engineered antibodies promote neutrophil-dependent control of Mycobacterium tuberculosis
- PMID: 39174703
- PMCID: PMC11371646
- DOI: 10.1038/s41564-024-01777-9
Fc-engineered antibodies promote neutrophil-dependent control of Mycobacterium tuberculosis
Abstract
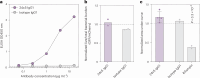
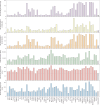
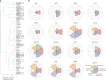
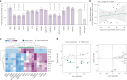
Mounting evidence indicates that antibodies can contribute towards control of tuberculosis (TB). However, the underlying mechanisms of humoral immune protection and whether antibodies can be exploited in therapeutic strategies to combat TB are relatively understudied. Here we engineered the receptor-binding Fc (fragment crystallizable) region of an antibody recognizing the Mycobacterium tuberculosis (Mtb) capsule, to define antibody Fc-mediated mechanism(s) of Mtb restriction. We generated 52 Fc variants that either promote or inhibit specific antibody effector functions, rationally building antibodies with enhanced capacity to promote Mtb restriction in a human whole-blood model of infection. While there is likely no singular Fc profile that universally drives control of Mtb, here we found that several Fc-engineered antibodies drove Mtb restriction in a neutrophil-dependent manner. Single-cell RNA sequencing analysis showed that a restrictive Fc-engineered antibody promoted neutrophil survival and expression of cell-intrinsic antimicrobial programs. These data show the potential of Fc-engineered antibodies as therapeutics able to harness the protective functions of neutrophils to promote control of TB.
© 2024. The Author(s).
Conflict of interest statement
G.A. is an employee of Moderna Therapeutics and holds equity in Leyden Labs and Seromyx Systems. M.Z.K. is an employee of Moderna Therapeutics. The other authors declare no competing interests.
Figures


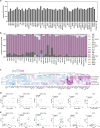




References
-
- Global Tuberculosis Report (World Health Organization, 2023).
MeSH terms
Substances
Grants and funding
- U54 CA225088/CA/NCI NIH HHS/United States
- U2CCA233280/Foundation for the National Institutes of Health (Foundation for the National Institutes of Health, Inc.)
- F31 AI150171/AI/NIAID NIH HHS/United States
- U2C CA233280/CA/NCI NIH HHS/United States
- U2C CA233262/CA/NCI NIH HHS/United States
- R01 AI166313/AI/NIAID NIH HHS/United States
- OPP1156795/Bill and Melinda Gates Foundation (Bill & Melinda Gates Foundation)
- U54CA225088/Foundation for the National Institutes of Health (Foundation for the National Institutes of Health, Inc.)
- R01 AI022553/AI/NIAID NIH HHS/United States
- U2CCA233262/Foundation for the National Institutes of Health (Foundation for the National Institutes of Health, Inc.)
- 75N93019C00071/AI/NIAID NIH HHS/United States
- AI150171-01/Foundation for the National Institutes of Health (Foundation for the National Institutes of Health, Inc.)
- R01A1022553/Foundation for the National Institutes of Health (Foundation for the National Institutes of Health, Inc.)
- P30 AI060354/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

