Computational investigations of potential inhibitors of monkeypox virus envelope protein E8 through molecular docking and molecular dynamics simulations
- PMID: 39179615
- PMCID: PMC11343748
- DOI: 10.1038/s41598-024-70433-3
Computational investigations of potential inhibitors of monkeypox virus envelope protein E8 through molecular docking and molecular dynamics simulations
Abstract
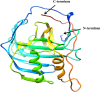
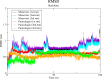
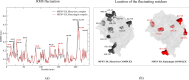
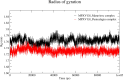
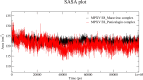
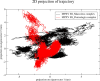
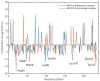
The World Health Organization (WHO) has declared the monkeypox outbreak a public health emergency, as there is no specific therapeutics for monkeypox virus (MPXV) disease. This study focused on docking various commercial drugs and plant-derived compounds against the E8 envelope protein crucial for MPXV attachment and pathogenesis. The target protein structure was modeled based on the vaccinia virus D8L protein. Notably, maraviroc and punicalagin emerged as potential ligands, with punicalagin exhibiting higher binding affinity (- 9.1 kcal/mol) than maraviroc (- 7.8 kcal/mol). Validation through 100 ns molecular dynamics (MD) simulations demonstrated increased stability of the E8-punicalagin complex, with lower RMSD, RMSF, and Rg compared to maraviroc. Enhanced hydrogen bonding, lower solvent accessibility, and compact motions also attributed to higher binding affinity and stability of the complex. MM-PBSA calculations revealed van der Waals, electrostatic, and non-polar solvation as principal stabilizing energies. The binding energy decomposition per residue favored stable interactions between punicalagin and the protein's active site residues (Arg20, Phe56, Glu228, Tyr232) compared to maraviroc. Overall study suggests that punicalagin can act as a potent inhibitor against MPXV. Further research and experimental investigations are warranted to validate its efficacy and safety.
Keywords: E8; MPXV; Maraviroc; Molecular docking; Molecular dynamics simulations; Punicalagin.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



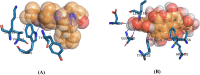



Similar articles
-
Machine learning and classical MD simulation to identify inhibitors against the P37 envelope protein of monkeypox virus.J Biomol Struct Dyn. 2024 May;42(8):3935-3948. doi: 10.1080/07391102.2023.2216290. Epub 2023 May 23. J Biomol Struct Dyn. 2024. PMID: 37221882
-
In-silico investigation of E8 surface protein of the monkeypox virus to identify potential therapeutic agents.J Biomol Struct Dyn. 2024 Oct;42(16):8242-8255. doi: 10.1080/07391102.2023.2245041. Epub 2023 Aug 9. J Biomol Struct Dyn. 2024. PMID: 37555596
-
Functional characterization and structural prediction of hypothetical proteins in monkeypox virus and identification of potential inhibitors.Mol Divers. 2025 Apr;29(2):1589-1617. doi: 10.1007/s11030-024-10935-4. Epub 2024 Jul 24. Mol Divers. 2025. PMID: 39043911
-
A review on computational studies and bioinformatics analysis of potential drugs against monkeypox virus.J Biomol Struct Dyn. 2024 Aug;42(12):6091-6107. doi: 10.1080/07391102.2023.2231542. Epub 2023 Jul 4. J Biomol Struct Dyn. 2024. PMID: 37403283 Review.
-
Discovery of Potent Inhibitors for the Inhibition of Dengue Envelope Protein: An In Silico Approach.Curr Top Med Chem. 2018;18(18):1585-1602. doi: 10.2174/1568026618666181025100736. Curr Top Med Chem. 2018. PMID: 30360716 Review.
Cited by
-
Structure-based drug designing for potential antiviral activity of selected natural product against Monkeypox (Mpox) virus and its host targets.Virusdisease. 2024 Dec;35(4):589-608. doi: 10.1007/s13337-024-00900-y. Epub 2024 Nov 29. Virusdisease. 2024. PMID: 39677842
-
Isolation and Characterization of E8 Monoclonal Antibodies from Donors Vaccinated with Recombinant Vaccinia Vaccine with Efficient Neutralization of Authentic Monkeypox Virus.Vaccines (Basel). 2025 Apr 27;13(5):471. doi: 10.3390/vaccines13050471. Vaccines (Basel). 2025. PMID: 40432083 Free PMC article.
-
Identification of potential VP40 inhibitor of Marburg virus through molecular docking, pharmacokinetic analysis and molecular dynamics simulation.Sci Rep. 2025 Jul 25;15(1):27129. doi: 10.1038/s41598-025-12917-4. Sci Rep. 2025. PMID: 40715601 Free PMC article.
-
Trans-Cannabitriol as a Dual Inhibition of MPOX Adhesion Receptors L1R and E8L: An In Silico Perspective.Bioinform Biol Insights. 2025 Jul 23;19:11779322251355315. doi: 10.1177/11779322251355315. eCollection 2025. Bioinform Biol Insights. 2025. PMID: 40718062 Free PMC article.
-
Virtual screening and identification of potent phytoconstituents from Acorus calamus L. as inhibitors of Monkeypox virus infection.J Genet Eng Biotechnol. 2025 Jun;23(2):100487. doi: 10.1016/j.jgeb.2025.100487. Epub 2025 Apr 25. J Genet Eng Biotechnol. 2025. PMID: 40390486 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

