Clinical and genomic diversity of Treponema pallidum subspecies pallidum to inform vaccine research: an international, molecular epidemiology study
- PMID: 39181152
- PMCID: PMC11371664
- DOI: 10.1016/S2666-5247(24)00087-9
Clinical and genomic diversity of Treponema pallidum subspecies pallidum to inform vaccine research: an international, molecular epidemiology study
Abstract
Background: The increase in syphilis rates worldwide necessitates development of a vaccine with global efficacy. We aimed to explore Treponema pallidum subspecies pallidum (TPA) molecular epidemiology essential for vaccine research by analysing clinical data and specimens from early syphilis patients using whole-genome sequencing (WGS) and publicly available WGS data.
Methods: In this multicentre, cross-sectional, molecular epidemiology study, we enrolled patients with primary, secondary, or early latent syphilis from clinics in China, Colombia, Malawi, and the USA between Nov 28, 2019, and May 27, 2022. Participants aged 18 years or older with laboratory confirmation of syphilis by direct detection methods or serological testing, or both, were included. Patients were excluded from enrolment if they were unwilling or unable to give informed consent, did not understand the study purpose or nature of their participation, or received antibiotics active against syphilis in the past 30 days. TPA detection and WGS were conducted on lesion swabs, skin biopsies, skin scrapings, whole blood, or rabbit-passaged isolates. We compared our WGS data to publicly available genomes and analysed TPA populations to identify mutations associated with lineage and geography.
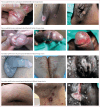
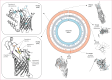
Findings: We screened 2802 patients and enrolled 233 participants, of whom 77 (33%) had primary syphilis, 154 (66%) had secondary syphilis, and two (1%) had early latent syphilis. The median age of participants was 28 years (IQR 22-35); 154 (66%) participants were cisgender men, 77 (33%) were cisgender women, and two (1%) were transgender women. Of the cisgender men, 66 (43%) identified as gay, bisexual, or other sexuality. Among all participants, 56 (24%) had HIV co-infection. WGS data from 113 participants showed a predominance of SS14-lineage strains with geographical clustering. Phylogenomic analyses confirmed that Nichols-lineage strains were more genetically diverse than SS14-lineage strains and clustered into more distinct subclades. Differences in single nucleotide variants (SNVs) were evident by TPA lineage and geography. Mapping of highly differentiated SNVs to three-dimensional protein models showed population-specific substitutions, some in outer membrane proteins (OMPs) of interest.
Interpretation: Our study substantiates the global diversity of TPA strains. Additional analyses to explore TPA OMP variability within strains is vital for vaccine development and understanding syphilis pathogenesis on a population level.
Funding: US National Institutes of Health National Institute for Allergy and Infectious Disease, the Bill & Melinda Gates Foundation, Connecticut Children's, and the Czech Republic National Institute of Virology and Bacteriology.
Copyright © 2024 The Author(s). Published by Elsevier Ltd.. All rights reserved.
Conflict of interest statement
Declaration of interests ACS reports royalties from UptoDate; honoraria from the University of Alabama at Birmingham; and support for meetings or travel from the American STD Association as a member of the Executive Board outside the scope of the current work. JAG-L reports honoraria from the Universidad de Antioquia; support for meetings or travel from Carnott Laboratories, Cantabria Labs, Epidermique, Pharmaderm, and Janssen; receipt of writing materials from Epidermique, Cantabria labs, Isdin, Pharmaderm, Skindrugs, Loreal, Galderma, Cetaphil, Cerave, Isispharma, Carnott, Janssen, Pharmalab, Novartis, Pfizer, and Lilly outside of the scope of work. JJJ reports membership in the Worldwide Antimalarial Resistance Network. KLH reports honoraria from the Eastern Virginia Medical School and the Lawrence Livermore National Laboratory. JDR receives royalties from Biokit, Chembio, and Span Diagnostics for syphilis serodiagnostic reagents; and support for meetings or travel from Indiana University outside the scope of the current work. JBP reports research support from Gilead Sciences; non-financial support from Abbott Diagnostics; and consulting for Zymeron Corporation, all outside the scope of the current work.
Figures



Update of
-
Clinical and genomic diversity of Treponema pallidum subsp. pallidum: A global, multi-center study of early syphilis to inform vaccine research.medRxiv [Preprint]. 2023 Jul 24:2023.07.19.23291250. doi: 10.1101/2023.07.19.23291250. medRxiv. 2023. Update in: Lancet Microbe. 2024 Sep;5(9):100871. doi: 10.1016/S2666-5247(24)00087-9. PMID: 37546832 Free PMC article. Updated. Preprint.
References
-
- Ghanem KG, Ram S, Rice PA. The modern epidemic of syphilis. N Engl J Med. 2020;382:845–854. - PubMed
-
- WHO Global progress report on HIV, viral hepatitis and sexually transmitted infections, 2021. Accountability for the global health sector strategies 2016–2021: actions for impact. 2021. https://www.who.int/publications/i/item/9789240027077
-
- Arora N, Schuenemann VJ, Jäger G, et al. Origin of modern syphilis and emergence of a pandemic Treponema pallidum cluster. Nat Microbiol. 2016;2 - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

