Spatiotemporal transcriptomic mapping of regenerative inflammation in skeletal muscle reveals a dynamic multilayered tissue architecture
- PMID: 39190487
- PMCID: PMC11473166
- DOI: 10.1172/JCI173858
Spatiotemporal transcriptomic mapping of regenerative inflammation in skeletal muscle reveals a dynamic multilayered tissue architecture
Abstract
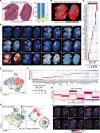
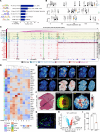
Tissue regeneration is orchestrated by macrophages that clear damaged cells and promote regenerative inflammation. How macrophages spatially adapt and diversify their functions to support the architectural requirements of actively regenerating tissue remains unknown. In this study, we reconstructed the dynamic trajectories of myeloid cells isolated from acutely injured and early stage dystrophic muscles. We identified divergent subsets of monocytes/macrophages and DCs and validated markers (e.g., glycoprotein NMB [GPNMB]) and transcriptional regulators associated with defined functional states. In dystrophic muscle, specialized repair-associated subsets exhibited distinct macrophage diversity and reduced DC heterogeneity. Integrating spatial transcriptomics analyses with immunofluorescence uncovered the ordered distribution of subpopulations and multilayered regenerative inflammation zones (RIZs) where distinct macrophage subsets are organized in functional zones around damaged myofibers supporting all phases of regeneration. Importantly, intermittent glucocorticoid treatment disrupted the RIZs. Our findings suggest that macrophage subtypes mediated the development of the highly ordered architecture of regenerative tissues, unveiling the principles of the structured yet dynamic nature of regenerative inflammation supporting effective tissue repair.
Keywords: Expression profiling; Inflammation; Macrophages; Skeletal muscle.
Figures






References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

