Single-Molecule Fingerprinting Reveals Different Growth Mechanisms in Seed Amplification Assays for Different Polymorphs of α-Synuclein Fibrils
- PMID: 39197832
- PMCID: PMC11413846
- DOI: 10.1021/acschemneuro.4c00185
Single-Molecule Fingerprinting Reveals Different Growth Mechanisms in Seed Amplification Assays for Different Polymorphs of α-Synuclein Fibrils
Abstract
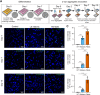
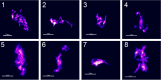
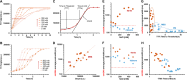
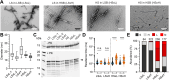
α-Synuclein (αSyn) aggregates, detected in the biofluids of patients with Parkinson's disease (PD), have the ability to catalyze their own aggregation, leading to an increase in the number and size of aggregates. This self-templated amplification is used by newly developed assays to diagnose Parkinson's disease and turns the presence of αSyn aggregates into a biomarker of the disease. It has become evident that αSyn can form fibrils with slightly different structures, called "strains" or polymorphs, but little is known about their differential reactivity in diagnostic assays. Here, we compared the properties of two well-described αSyn polymorphs. Using single-molecule techniques, we observed that one of the polymorphs had an increased tendency to undergo secondary nucleation and we showed that this could explain the differences in reactivity observed in in vitro seed amplification assay and cellular assays. Simulations and high-resolution microscopy suggest that a 100-fold difference in the apparent rate of growth can be generated by a surprisingly low number of secondary nucleation "points" (1 every 2000 monomers added by elongation). When both strains are present in the same seeded reaction, secondary nucleation displaces proportions dramatically and causes a single strain to dominate the reaction as the major end product.
Keywords: PMCA; Parkinson’s disease; RT-QuIC; preformed fibrils; seed amplification assay; single-molecule detection; strains; α-synuclein.
Conflict of interest statement
The authors declare the following competing financial interest(s): ES and YG are the inventors of the AttoBright instrument, which is patented; ES and YG are the founders of AttoQuest Pty Ltd, an official spin-off from UNSW that commercializes the AttoBright instruments.
Figures




Similar articles
-
Single Molecule Fingerprinting Reveals Different Amplification Properties of α-Synuclein Oligomers and Preformed Fibrils in Seeding Assay.ACS Chem Neurosci. 2022 Apr 6;13(7):883-896. doi: 10.1021/acschemneuro.1c00553. Epub 2022 Mar 14. ACS Chem Neurosci. 2022. PMID: 35286811 Free PMC article.
-
The Alpha-Synuclein RT-QuIC Products Generated by the Olfactory Mucosa of Patients with Parkinson's Disease and Multiple System Atrophy Induce Inflammatory Responses in SH-SY5Y Cells.Cells. 2021 Dec 28;11(1):87. doi: 10.3390/cells11010087. Cells. 2021. PMID: 35011649 Free PMC article.
-
α-Synuclein Seeding Assay Using RT-QuIC.Methods Mol Biol. 2021;2322:3-16. doi: 10.1007/978-1-0716-1495-2_1. Methods Mol Biol. 2021. PMID: 34043187
-
RT-QuIC and Related Assays for Detecting and Quantifying Prion-like Pathological Seeds of α-Synuclein.Biomolecules. 2022 Apr 14;12(4):576. doi: 10.3390/biom12040576. Biomolecules. 2022. PMID: 35454165 Free PMC article. Review.
-
α-Synuclein Strains and Their Relevance to Parkinson's Disease, Multiple System Atrophy, and Dementia with Lewy Bodies.Int J Mol Sci. 2023 Jul 28;24(15):12134. doi: 10.3390/ijms241512134. Int J Mol Sci. 2023. PMID: 37569510 Free PMC article. Review.
Cited by
-
Dementia with Lewy bodies and Parkinson disease dementia - the same or different and is it important?Nat Rev Neurol. 2025 Jul;21(7):394-403. doi: 10.1038/s41582-025-01090-x. Epub 2025 May 12. Nat Rev Neurol. 2025. PMID: 40355531 Review.
References
-
- Van der Perren A.; Gelders G.; Fenyi A.; Bousset L.; Brito F.; Peelaerts W.; Van den Haute C.; Gentleman S.; Melki R.; Baekelandt V. The structural differences between patient-derived α-synuclein strains dictate characteristics of Parkinson’s disease, multiple system atrophy and dementia with Lewy bodies. Acta Neuropathol. 2020, 139 (6), 977–1000. 10.1007/s00401-020-02157-3. - DOI - PMC - PubMed
-
- Buell A. K.; Galvagnion C.; Gaspar R.; Sparr E.; Vendruscolo M.; Knowles T. P.; Linse S.; Dobson C. M. Solution conditions determine the relative importance of nucleation and growth processes in α-synuclein aggregation. Proc. Natl. Acad. Sci. U.S.A. 2014, 111 (21), 7671–7676. 10.1073/pnas.1315346111. - DOI - PMC - PubMed
-
- Russo M. J.; Orru C. D.; Concha-Marambio L.; Giaisi S.; Groveman B. R.; Farris C. M.; Holguin B.; Hughson A. G.; LaFontant D. E.; Caspell-Garcia C.; Coffey C. S.; Mollon J.; Hutten S. J.; Merchant K.; Heym R. G.; Soto C.; Caughey B.; Kang U. J. High diagnostic performance of independent alpha-synuclein seed amplification assays for detection of early Parkinson’s disease. Acta Neuropathol. Commun. 2021, 9 (1), 17910.1186/s40478-021-01282-8. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

