Community Resource: Large-Scale Proteogenomics to Refine Wheat Genome Annotations
- PMID: 39201310
- PMCID: PMC11354340
- DOI: 10.3390/ijms25168614
Community Resource: Large-Scale Proteogenomics to Refine Wheat Genome Annotations
Abstract
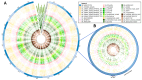
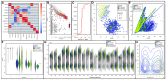
Triticum aestivum is an important crop whose reference genome (International Wheat Genome Sequencing Consortium (IWGSC) RefSeq v2.1) offers a valuable resource for understanding wheat genetic structure, improving agronomic traits, and developing new cultivars. A key aspect of gene model annotation is protein-level evidence of gene expression obtained from proteomics studies, followed up by proteogenomics to physically map proteins to the genome. In this research, we have retrieved the largest recent wheat proteomics datasets publicly available and applied the Basic Local Alignment Search Tool (tBLASTn) algorithm to map the 861,759 identified unique peptides against IWGSC RefSeq v2.1. Of the 92,719 hits, 83,015 unique peptides aligned along 33,612 High Confidence (HC) genes, thus validating 31.4% of all wheat HC gene models. Furthermore, 6685 unique peptides were mapped against 3702 Low Confidence (LC) gene models, and we argue that these gene models should be considered for HC status. The remaining 2934 orphan peptides can be used for novel gene discovery, as exemplified here on chromosome 4D. We demonstrated that tBLASTn could not map peptides exhibiting mid-sequence frame shift. We supply all our proteogenomics results, Galaxy workflow and Python code, as well as Browser Extensible Data (BED) files as a resource for the wheat community via the Apollo Jbrowse, and GitHub repositories. Our workflow could be applied to other proteomics datasets to expand this resource with proteins and peptides from biotically and abiotically stressed samples. This would help tease out wheat gene expression under various environmental conditions, both spatially and temporally.
Keywords: Triticum aestivum; bottom-up proteogenomics; gene models; genome annotation; multiple sequence alignment; proteomics.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures



References
-
- The International Wheat Genome Sequencing Consortium (IWGSC) Appels R., Eversole K., Stein N., Feuillet C., Keller B., Rogers J., Pozniak C.J., Choulet F., Distelfeld A., et al. Shifting the Limits in Wheat Research and Breeding Using a Fully Annotated Reference Genome. Science. 2018;361:eaar7191. doi: 10.1126/science.aar7191. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

