Role of Extracellular Vesicles in Crohn's Patients on Adalimumab Who Received COVID-19 Vaccination
- PMID: 39201543
- PMCID: PMC11355036
- DOI: 10.3390/ijms25168853
Role of Extracellular Vesicles in Crohn's Patients on Adalimumab Who Received COVID-19 Vaccination
Abstract
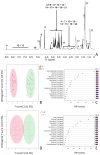
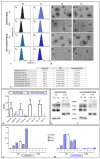
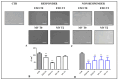
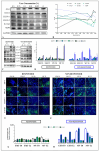
Crohn's disease (CD) is a type of inflammatory bowel disease (IBD) affecting the gastrointestinal tract that can also cause extra-intestinal complications. Following exposure to the mRNA vaccine BNT162b2 (Pfizer-BioNTech) encoding the SARS-CoV-2 Spike (S) protein, some patients experienced a lack of response to the biological drug Adalimumab and a recrudescence of the disease. In CD patients in progression, resistant to considered biological therapy, an abnormal increase in intestinal permeability was observed, more often with a modulated expression of different proteins such as Aquaporin 8 (AQP8) and in tight junctions (e.g., ZO-1, Claudin1, Claudin2, Occludin), especially during disease flares. The aim of this study is to investigate how the SARS-CoV-2 vaccine could interfere with IBD therapy and contribute to disease exacerbation. We investigated the role of the SARS-CoV-2 Spike protein, transported by extracellular vesicles (EVs), and the impact of various EVs components, namely, exosomes (EXOs) and microvesicles (MVs), in modulating the expression of molecules involved in the exacerbation of CD, which remains unknown.
Keywords: Adalimumab; Crohn’s disease; SARS-Cov-2; exosomes; ions channel; microvesicles.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

References
-
- Coward S., Benchimol E.I., Kuenzig M.E., Windsor J.W., Bernstein C.N., Bitton A., Jones J.L., Lee K., Murthy S.K., Targownik L.E., et al. The 2023 Impact of Inflammatory Bowel Disease in Canada: Epidemiology of IBD. J. Can. Assoc. Gastroenterol. 2023;6:256. doi: 10.1093/jcag/gwad004. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

