Exploring Mitochondrial Heterogeneity and Evolutionary Dynamics in Thelephora ganbajun through Population Genomics
- PMID: 39201699
- PMCID: PMC11354633
- DOI: 10.3390/ijms25169013
Exploring Mitochondrial Heterogeneity and Evolutionary Dynamics in Thelephora ganbajun through Population Genomics
Abstract
Limited exploration in fungal mitochondrial genetics has uncovered diverse inheritance modes. The mitochondrial genomes are inherited uniparentally in the majority of sexual eukaryotes, our discovery of persistent mitochondrial heterogeneity within the natural population of the basidiomycete fungus Thelephora ganbajun represents a significant advance in understanding mitochondrial inheritance and evolution in eukaryotes. Here, we present a comprehensive analysis by sequencing and assembling the complete mitogenomes of 40 samples exhibiting diverse cox1 heterogeneity patterns from various geographical origins. Additionally, we identified heterogeneous variants in the nad5 gene, which, similar to cox1, displayed variability across multiple copies. Notably, our study reveals a distinct prevalence of introns and homing endonucleases in these heterogeneous genes. Furthermore, we detected potential instances of horizontal gene transfer involving homing endonucleases. Population genomic analyses underscore regional variations in mitochondrial genome composition among natural samples exhibiting heterogeneity. Thus, polymorphisms in heterogeneous genes, introns, and homing endonucleases significantly influence mitochondrial structure, structural variation, and evolutionary dynamics in this species. This study contributes valuable insights into mitochondrial genome architecture, population dynamics, and the evolutionary implications of mitochondrial heterogeneity in sexual eukaryotes.
Keywords: comparative population genomics; mitogenome; persistent heteroplasmy; wild edible fungi.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

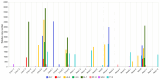
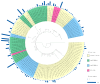
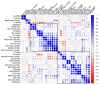

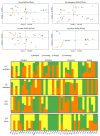
References
-
- Araújo D.S., De-Paula R.B., Tomé L.M.R., Quintanilha-Peixoto G., Salvador-Montoya C.A., Del-Bem L.-E., Badotti F., Azevedo V.A.C., Brenig B., Aguiar E.R.G.R., et al. Comparative mitogenomics of Agaricomycetes: Diversity, abundance, impact and coding potential of putative open-reading frames. Mitochondrion. 2021;58:1–13. doi: 10.1016/j.mito.2021.02.002. - DOI - PubMed
-
- Liu W., Cai Y., Zhang Q., Chen L., Shu F., Ma X., Bian Y. The mitochondrial genome of Morchella importuna (272.2 kb) is the largest among fungi and contains numerous introns, mitochondrial non-conserved open reading frames and repetitive sequences. Int. J. Biol. Macromol. 2020;143:373–381. doi: 10.1016/j.ijbiomac.2019.12.056. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

