Identification and Prediction of Differentially Expressed MicroRNAs Associated with Detoxification Pathways in Larvae of Spodoptera frugiperda
- PMID: 39202382
- PMCID: PMC11353827
- DOI: 10.3390/genes15081021
Identification and Prediction of Differentially Expressed MicroRNAs Associated with Detoxification Pathways in Larvae of Spodoptera frugiperda
Abstract
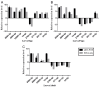
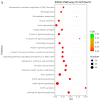
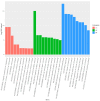
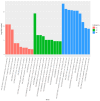
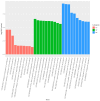
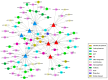
Spodoptera frugiperda poses a severe threat to crops, causing substantial economic losses. The increased use of chemical pesticides has led to resistance in S. frugiperda populations. Micro ribonucleic acids (MicroRNAs or miRNAs) are pivotal in insect growth and development. This study aims to identify miRNAs across different developmental stages of S. frugiperda to explore differential expression and predict target gene functions. High-throughput sequencing of miRNAs was conducted on eggs, 3rd instar larvae, pupae, and adults. Bioinformatics analyses identified differentially expressed miRNAs specifically in larvae, with candidate miRNAs screened to predict target genes, particularly those involved in detoxification pathways. A total of 184 known miRNAs and 209 novel miRNAs were identified across stages. Comparative analysis revealed 54, 15, and 18 miRNAs differentially expressed in larvae, compared to egg, pupa, and adult stages, respectively. Eight miRNAs showed significant differential expression across stages, validated by quantitative reverse transcription PCR (qRT-PCR). Gene Ontology and Kyoto Encyclopedia of Genes and Genomes enrichment analyses predicted target genes' functions, identifying eight differentially expressed miRNAs targeting 10 gene families associated with detoxification metabolism, including P450s, glutathione S-transferase (GSTs), ATP-binding cassette (ABC) transporters, and sodium channels. These findings elucidate the species-specific miRNA profiles and regulatory mechanisms of detoxification-related genes in S. frugiperda larvae, offering insights and strategies for effectively managing this pest.
Keywords: S. frugiperda; detoxification; larva; microRNAs; resistance; the fall armyworm.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures


Similar articles
-
Transcriptomics and metagenomics of common cutworm (Spodoptera litura) and fall armyworm (Spodoptera frugiperda) demonstrate differences in detoxification and development.BMC Genomics. 2022 May 20;23(1):388. doi: 10.1186/s12864-022-08613-6. BMC Genomics. 2022. PMID: 35596140 Free PMC article.
-
Differential detoxification enzyme profiles in C-corn strain and R-rice strain of Spodoptera frugiperda by comparative genomic analysis: insights into host adaptation.BMC Genomics. 2025 Jan 6;26(1):14. doi: 10.1186/s12864-024-11185-2. BMC Genomics. 2025. PMID: 39762739 Free PMC article.
-
Comprehensive transcriptome and metabolome analysis of the adaptability and detoxification ability of Spodoptera frugiperda larvae to tobacco.J Insect Physiol. 2025 Jun;163:104800. doi: 10.1016/j.jinsphys.2025.104800. Epub 2025 Apr 11. J Insect Physiol. 2025. PMID: 40221127
-
Role of glutathione S-transferase gene SfGSTs2 in the host plant adaptation of the polyphagous pest Spodoptera frugiperda.Bull Entomol Res. 2025 Aug;115(4):524-535. doi: 10.1017/S0007485325100084. Bull Entomol Res. 2025. PMID: 40509801
-
Comparative analysis of the detoxification gene inventory of four major Spodoptera pest species in response to xenobiotics.Insect Biochem Mol Biol. 2021 Nov;138:103646. doi: 10.1016/j.ibmb.2021.103646. Epub 2021 Aug 29. Insect Biochem Mol Biol. 2021. PMID: 34469782 Review.
Cited by
-
Developmental Expression Patterns of miRNA in Mythimna separata Walker (Lepidotera: Noctuidae).Genes (Basel). 2025 Feb 19;16(2):234. doi: 10.3390/genes16020234. Genes (Basel). 2025. PMID: 40004562 Free PMC article.
References
MeSH terms
Substances
Grants and funding
- 2023NSFSC1258/Identification and functional prediction of preferred expression microRNAs in Spodoptera frugi-perda larvae
- 2023NSFSC0152/Research on the mechanism of the identification of Telenomus remus Nixon and the parasitism of Spodoptera frugiperda
- 2021XKJS084/Study on the disaster law and key control technology of main pests in Sichuan characteristic fruits
- sccxtd-04/Sichuan Fruit Innovation Team of National Modern Agricultural Industry Technology System
LinkOut - more resources
Full Text Sources

