Integration of Bioinformatics and Machine Learning to Identify CD8+ T Cell-Related Prognostic Signature to Predict Clinical Outcomes and Treatment Response in Breast Cancer Patients
- PMID: 39202452
- PMCID: PMC11353403
- DOI: 10.3390/genes15081093
Integration of Bioinformatics and Machine Learning to Identify CD8+ T Cell-Related Prognostic Signature to Predict Clinical Outcomes and Treatment Response in Breast Cancer Patients
Abstract
The incidence of breast cancer (BC) continues to rise steadily, posing a significant burden on the public health systems of various countries worldwide. As a member of the tumor microenvironment (TME), CD8+ T cells inhibit cancer progression through their protective role. This study aims to investigate the role of CD8+ T cell-related genes (CTRGs) in breast cancer patients.
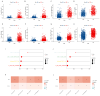
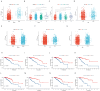
Methods: We assessed the abundance of CD8+ T cells in the TCGA and METABRIC datasets and obtained CTRGs through WGCNA. Subsequently, a prognostic signature (CTR score) was constructed from CTRGs screened by seven machine learning algorithms, and the relationship between the CTR score and TME, immunotherapy, and drug sensitivity was analyzed. Additionally, CTRGs' expression in different cells within TME was identified through single-cell analysis and spatial transcriptomics. Finally, the expression of CTRGs in clinical tissues was verified via RT-PCR.
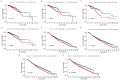
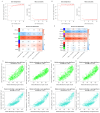
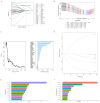
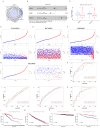
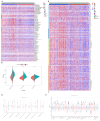
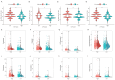
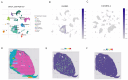
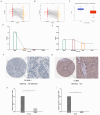
Results: The CD8+ T cell-related prognostic signature consists of two CTRGs. In the TCGA and METABRIC datasets, the CTR score appeared to be negatively linked to the abundance of CD8+ T cells, and BC patients with higher risk score show a worse prognosis. The low CTR score group exhibits higher immune infiltration levels, closely associated with inhibiting the tumor microenvironment. Compared with the high CTR score group, the low CTR score group shows better responses to chemotherapy and immune checkpoint therapy. Single-cell analysis and spatial transcriptomics reveal the heterogeneity of two CTRGs in different cells. Compared with the adjacent tissues, CD163L1 and KLRB1 mRNA are downregulated in tumor tissues.
Conclusions: This study establishes a robust CD8+ T cell-related prognostic signature, providing new insights for predicting the clinical outcomes and treatment responses of breast cancer patients.
Keywords: CD8+ T cell; breast cancer; machine learning; prognostic signature; tumor microenvironment.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

References
-
- Loibl S., André F., Bachelot T., Barrios C.H., Bergh J., Burstein H.J., Cardoso M.J., Carey L.A., Dawood S., Del Mastro L., et al. Early Breast Cancer: ESMO Clinical Practice Guideline for Diagnosis, Treatment and Follow-Up. Ann. Oncol. 2024;35:159–182. doi: 10.1016/j.annonc.2023.11.016. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

