Local Ancestry Inference Based on Population-Specific Single-Nucleotide Polymorphisms-A Study of Admixed Populations in the 1000 Genomes Project
- PMID: 39202458
- PMCID: PMC11353365
- DOI: 10.3390/genes15081099
Local Ancestry Inference Based on Population-Specific Single-Nucleotide Polymorphisms-A Study of Admixed Populations in the 1000 Genomes Project
Abstract
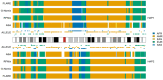
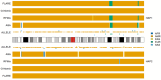
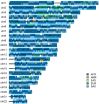
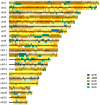
Human populations have interacted throughout history, and a considerable portion of modern human populations show evidence of admixture. Local ancestry inference (LAI) is focused on detecting the genetic ancestry of chromosomal segments in admixed individuals and has wide applications. In this work, we proposed a new LAI method based on population-specific single-nucleotide polymorphisms (SNPs) and applied it in the analysis of admixed populations in the 1000 Genomes Project (1KGP). Based on population-specific SNPs in a sliding window, we computed local ancestry information vectors, which are moment estimators of local ancestral proportions, for two haplotypes of an admixed individual and inferred the local ancestral origins. Then we used African (AFR), East Asian (EAS), European (EUR) and South Asian (SAS) populations from the 1KGP and indigenous American (AMR) populations from the Human Genome Diversity Project (HGDP) as reference populations and conducted the proposed LAI analysis on African American populations and American populations in the 1KGP. The results were compared with those obtained by RFMix, G-Nomix and FLARE. We demonstrated that the existence of alleles in a chromosomal region that are specific to a particular reference population and the absence of alleles specific to the other reference populations provide reasonable evidence for determining the ancestral origin of the region. Contemporary AFR, AMR and EUR populations approximate ancestral populations of the admixed populations well, and the results from RFMix, G-Nomix and FLARE largely agree with those from the Ancestral Spectrum Analyzer (ASA), in which the proposed method was implemented. When admixtures are ancient and contemporary reference populations do not satisfactorily approximate ancestral populations, the performances of RFMix, G-Nomix and FLARE deteriorate with increased error rates and fragmented chromosomal segments. In contrast, our method provides fair results.
Keywords: admixture; human populations; local ancestry inference; population-specific SNPs.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures





Similar articles
-
Opportunities and challenges of local ancestry in genetic association analyses.Am J Hum Genet. 2025 Apr 3;112(4):727-740. doi: 10.1016/j.ajhg.2025.03.004. Am J Hum Genet. 2025. PMID: 40185073 Review.
-
Characterizing features affecting local ancestry inference performance in admixed populations.Am J Hum Genet. 2025 Feb 6;112(2):224-234. doi: 10.1016/j.ajhg.2024.12.005. Epub 2025 Jan 2. Am J Hum Genet. 2025. PMID: 39753130 Free PMC article.
-
Distribution and linkage disequilibrium of the enhancer SNP rs5758550 among Latin American populations: influence of continental ancestry.Pharmacogenet Genomics. 2020 Jun;30(4):67-72. doi: 10.1097/FPC.0000000000000398. Pharmacogenet Genomics. 2020. PMID: 32187157
-
An ancestry informative marker panel design for individual ancestry estimation of Hispanic population using whole exome sequencing data.BMC Genomics. 2019 Dec 30;20(Suppl 12):1007. doi: 10.1186/s12864-019-6333-6. BMC Genomics. 2019. PMID: 31888480 Free PMC article.
-
The genomic landscape of African populations in health and disease.Hum Mol Genet. 2017 Oct 1;26(R2):R225-R236. doi: 10.1093/hmg/ddx253. Hum Mol Genet. 2017. PMID: 28977439 Free PMC article. Review.
Cited by
-
Genome-wide association analyses reveal susceptibility variants linked to Parkinson's disease in the South African population using inferred global and local ancestry.medRxiv [Preprint]. 2025 Aug 2:2025.08.01.25331910. doi: 10.1101/2025.08.01.25331910. medRxiv. 2025. PMID: 40766124 Free PMC article. Preprint.
-
Optimization of computational ancestry inference for use in cancer cell lines.Biol Methods Protoc. 2025 Jun 2;10(1):bpaf043. doi: 10.1093/biomethods/bpaf043. eCollection 2025. Biol Methods Protoc. 2025. PMID: 40585183 Free PMC article.
-
Opportunities and challenges of local ancestry in genetic association analyses.Am J Hum Genet. 2025 Apr 3;112(4):727-740. doi: 10.1016/j.ajhg.2025.03.004. Am J Hum Genet. 2025. PMID: 40185073 Review.
References
MeSH terms
LinkOut - more resources
Full Text Sources

