Prevalence of Respiratory Pathogens in Nasopharyngeal Swabs of Febrile Patients with or without Respiratory Symptoms in the Niakhar Area of Rural Senegal
- PMID: 39204255
- PMCID: PMC11357141
- DOI: 10.3390/pathogens13080655
Prevalence of Respiratory Pathogens in Nasopharyngeal Swabs of Febrile Patients with or without Respiratory Symptoms in the Niakhar Area of Rural Senegal
Abstract
Acute respiratory tract infections are one of the leading causes of morbidity and mortality worldwide. More data are needed on circulating respiratory microorganisms in different geographical areas and ecosystems. We analyzed nasopharyngeal swabs from 500 febrile patients living in the Niakhar area (Senegal), using FTDTM multiplex qPCR and simplex qPCR to target a panel of 25 microorganisms. We detected at least one microorganism for 366/500 patients (73.2%), at least one virus for 193/500 (38.6%), and at least one bacterium for 324/500 (64.8%). The most frequently detected microorganisms were Streptococcus pneumoniae (36.8%), Haemophilus influenzae (35.8%), adenovirus (11.8%), influenza viruses (6.4%), rhinovirus (5.0%), SARS-CoV-2 (4.0%), and RSV (4.0%). The main microorganisms significantly associated with respiratory symptoms, with a p-value ≤ 0.05, were influenza virus (11.9% in patients with respiratory symptoms versus 2.9% in patients without), RSV (6.5% versus 2.6%), metapneumovirus (5.4% versus 1.3%), HPIVs (7.6% versus 1.0%), S. pneumoniae (51.9% versus 28.0%), and H. influenzae (54.6% versus 24.5%). Co-infections were significantly associated with respiratory symptoms (65.4% versus 32.9%). All the epidemiological data show a high level of circulation of respiratory pathogens among febrile patients, including those preventable by vaccination such as S. pneumoniae, raising the question of the serotypes currently circulating. Furthermore, the availability of affordable real-time etiological diagnostic tools would enable management to be adapted as effectively as possible.
Keywords: Haemophilus influenzae; RSV; SARS-CoV-2; Senegal; Streptococcus pneumoniae; acute respiratory tract infections; adenovirus; human coronaviruses; influenza; sub-Saharan Africa.
Conflict of interest statement
The authors have no conflicts of interest to declare. Funding sources played no role in the design and conduct of the study; collection, management, analysis, and interpretation of the data; and preparation, review, or approval of the manuscript.
Figures

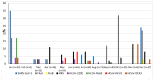
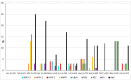

References
-
- van Rijn A.L., Nijhuis R.H.T., Bekker V., Groeneveld G.H., Wessels E., Feltkamp M.C.W., Claas E.C.J. Clinical implications of rapid ePlex® respiratory pathogen panel testing compared to laboratory-developed real-time PCR. Eur. J. Clin. Microbiol. Infect. Dis. 2018;37:571–577. doi: 10.1007/s10096-017-3151-0. - DOI - PMC - PubMed
-
- Dare R., Sanghavi S., Bullotta A., Keightley M.-C., George K.S., Wadowsky R.M., Paterson D.L., McCurry K.R., Reinhart T.A., Husain S., et al. Diagnosis of human metapneumovirus infection in immunosuppressed lung transplant recipients and children evaluated for pertussis. J. Clin. Microbiol. 2007;45:548–552. doi: 10.1128/JCM.01621-06. - DOI - PMC - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

