The global population stru cture of Lacticaseibacillus rhamnosus and its application to an investigation of a rare case of infective endocarditis
- PMID: 39213326
- PMCID: PMC11364288
- DOI: 10.1371/journal.pone.0300843
The global population stru cture of Lacticaseibacillus rhamnosus and its application to an investigation of a rare case of infective endocarditis
Abstract
Background: Lacticaseibacillus (formerly Lactobacillus) rhamnosus is widely used in probiotics or food supplements to promote microbiome health and may also be part of the normal microbiota of the human gastrointestinal tract. However, it rarely also causes invasive or severe infections in patients. It has been postulated that these infections may originate from probiotics or from endogenous commensal reservoirs. In this report, we examine the population structure of Lacticaseibacillus rhamnosus and investigate the utility of using bacterial genomics to identify the source of invasive Lacticaseibacillus infections.
Methods: Core genome phylogenetic analysis was performed on 602 L. rhamnosus genome sequences from the National Center for Biotechnology public database. This information was then used along with newly generated sequences of L. rhamnosus isolates from yogurt to investigate a fatal case of L. rhamnosus endocarditis.
Results: Phylogenetic analysis demonstrated substantial genetic overlap of L. rhamnosus isolates cultured from food, probiotics, infected patients, and colonized individuals. This was applied to a patient who had both consumed yogurt and developed L. rhamnosus endocarditis to attempt to identify the source of his infection. The sequence of the isolate from the patient's bloodstream differed at only one nucleotide position from one of the yogurt isolates. Both isolates belonged to a clade, identified here as clade YC, composed of mostly gastrointestinal isolates from healthy individuals, some of which also differed by only a single nucleotide change from the patient's isolate.
Conclusions: As illustrated by this case, whole genome sequencing may be insufficient to reliably determine the source of invasive infections caused by L. rhamnosus.
Copyright: © 2024 Santoiemma et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

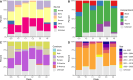

References
-
- Zheng J, Wittouck S, Salvetti E, Franz C, Harris HMB, Mattarelli P, et al. A taxonomic note on the genus Lactobacillus: Description of 23 novel genera, emended description of the genus Lactobacillus Beijerinck 1901, and union of Lactobacillaceae and Leuconostocaceae. Int J Syst Evol Microbiol. 2020;70(4):2782–858. Epub 2020/04/16. doi: 10.1099/ijsem.0.004107 . - DOI - PubMed
-
- US Food & Drug Administration. GRN No. 1093 Lacticaseibacillus rhamnosus IDCC 3201. 2023. Available from: https://www.cfsanappsexternal.fda.gov/scripts/fdcc/?set=GRASNotices&id=1093.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

