AI-powered omics-based drug pair discovery for pyroptosis therapy targeting triple-negative breast cancer
- PMID: 39215014
- PMCID: PMC11364624
- DOI: 10.1038/s41467-024-51980-9
AI-powered omics-based drug pair discovery for pyroptosis therapy targeting triple-negative breast cancer
Abstract
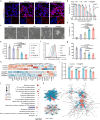
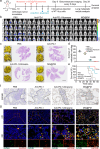
Due to low success rates and long cycles of traditional drug development, the clinical tendency is to apply omics techniques to reveal patient-level disease characteristics and individualized responses to treatment. However, the heterogeneous form of data and uneven distribution of targets make drug discovery and precision medicine a non-trivial task. This study takes pyroptosis therapy for triple-negative breast cancer (TNBC) as a paradigm and uses data mining of a large TNBC cohort and drug databases to establish a biofactor-regulated neural network for rapidly screening and optimizing compound pyroptosis drug pairs. Subsequently, biomimetic nanococrystals are prepared using the preferred combination of mitoxantrone and gambogic acid for rational drug delivery. The unique mechanism of obtained nanococrystals regulating pyroptosis genes through ribosomal stress and triggering pyroptosis cascade immune effects are revealed in TNBC models. In this work, a target omics-based intelligent compound drug discovery framework explores an innovative drug development paradigm, which repurposes existing drugs and enables precise treatment of refractory diseases.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





References
-
- Gandomi, A. & Haider, M. Beyond the hype: big data concepts, methods, and analytics. Int. J. Inf. Manag.35, 137–144 (2015). 10.1016/j.ijinfomgt.2014.10.007 - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

