LUBAC enables tumor-promoting LTβ receptor signaling by activating canonical NF-κB
- PMID: 39215104
- PMCID: PMC11445442
- DOI: 10.1038/s41418-024-01355-w
LUBAC enables tumor-promoting LTβ receptor signaling by activating canonical NF-κB
Abstract
Lymphotoxin β receptor (LTβR), a member of the TNF receptor superfamily (TNFR-SF), is essential for development and maturation of lymphoid organs. In addition, LTβR activation promotes carcinogenesis by inducing a proinflammatory secretome. Yet, we currently lack a detailed understanding of LTβR signaling. In this study we discovered the linear ubiquitin chain assembly complex (LUBAC) as a previously unrecognized and functionally crucial component of the native LTβR signaling complex (LTβR-SC). Mechanistically, LUBAC-generated linear ubiquitin chains enable recruitment of NEMO, OPTN and A20 to the LTβR-SC, where they act coordinately to regulate the balance between canonical and non-canonical NF-κB pathways. Thus, different from death receptor signaling, where LUBAC prevents inflammation through inhibition of cell death, in LTβR signaling LUBAC is required for inflammatory signaling by enabling canonical and interfering with non-canonical NF-κB activation. This results in a LUBAC-dependent LTβR-driven inflammatory, protumorigenic secretome. Intriguingly, in liver cancer patients with high LTβR expression, high expression of LUBAC correlates with poor prognosis, providing clinical relevance for LUBAC-mediated inflammatory LTβR signaling.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



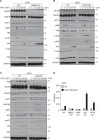


References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

