USP36 promotes tumorigenesis and tamoxifen resistance in breast cancer by deubiquitinating and stabilizing ERα
- PMID: 39215346
- PMCID: PMC11365244
- DOI: 10.1186/s13046-024-03160-2
USP36 promotes tumorigenesis and tamoxifen resistance in breast cancer by deubiquitinating and stabilizing ERα
Abstract
Background: Breast cancer is the most prevalent cancer in women globally. Over-activated estrogen receptor (ER) α signaling is considered the main factor in luminal breast cancers, which can be effectively managed with selective estrogen receptor modulators (SERMs) like tamoxifen. However, approximately 30-40% of ER + breast cancer cases are recurrent after tamoxifen therapy. This implies that the treatment of breast cancer is still hindered by resistance to tamoxifen. Recent studies have suggested that post-translational modifications of ERα play a significant role in endocrine resistance. The stability of both ERα protein and its transcriptome is regulated by a balance between E3 ubiquitin ligases and deubiquitinases. According to the current knowledge, approximately 100 deubiquitinases are encoded in the human genome, but it remains unclear which deubiquitinases play a critical role in estrogen signaling and endocrine resistance. Thus, decoding the key deubiquitinases that significantly impact estrogen signaling, including the control of ERα expression and stability, is critical for the improvement of breast cancer therapeutics.
Methods: We used several ER positive breast cancer cell lines, DUB siRNA library screening, xenograft models, endocrine-resistant (ERα-Y537S) model and performed immunoblotting, real time PCR, RNA sequencing, immunofluorescence, and luciferase activity assay to investigate the function of USP36 in breast cancer progression and tamoxifen resistance.
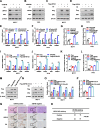
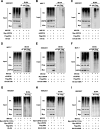
Results: In this study, we identify Ubiquitin-specific peptidase 36 (USP36) as a key deubiquitinase involved in ERα signaling and the advancement of breast cancer by deubiquitinases siRNA library screening. In vitro and in vivo studies showed that USP36, but not its catalytically inactive mutant (C131A), could promote breast cancer progression through ERα signaling. Conversely, silencing USP36 inhibited tumorigenesis. In models resistant to endocrine therapy, silencing USP36 destabilized the resistant form of ERα (Y537S) and restored sensitivity to tamoxifen. Molecular studies indicated that USP36 inhibited K48-linked polyubiquitination of ERα and enhanced the ERα transcriptome. It is interesting to note that our results suggest USP36 as a novel biomarker for treatment of breast cancer.
Conclusion: Our study revealed the possibility that inhibiting USP36 combined with tamoxifen could provide a potential therapy for breast cancer.
Keywords: Breast cancer; ERα; Stability; Tamoxifen resistance; USP36; Ubiquitin.
© 2024. The Author(s).
Conflict of interest statement
There is no competing interesting to declaim.
Figures







References
MeSH terms
Substances
Grants and funding
- No. 82203507/National Natural Science Foundation of China
- No. 82172999/National Natural Science Foundation of China
- No.82372969/National Natural Science Foundation of China
- Grant No. 21HASTIT049/Science and Technology Innovation Talents in Universities of Henan Province
- 222300420065/Natural Science Foundation of Henan Province
- No. 232102310291/Henan Province Science and Technology Innovation Talent Program
- tsqn202103175/Taishan Scholar Foundation of Shandong Province
- tsqn202306365/Taishan Scholar Foundation of Shandong Province
- ZR2021MH017/Natural Science Foundation of Shandong Province
- 22A310005/Foundation of Henan Educational Committee
- 005/University of Science and Technology Liaoning
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

