This is a preprint.
HMGB1 restores a dynamic chromatin environment in the presence of linker histone by deforming nucleosomal DNA
- PMID: 39229246
- PMCID: PMC11370580
- DOI: 10.1101/2024.08.23.609244
HMGB1 restores a dynamic chromatin environment in the presence of linker histone by deforming nucleosomal DNA
Update in
-
HMGB1 deforms nucleosomal DNA to generate a dynamic chromatin environment counteracting the effects of linker histone.Sci Adv. 2025 Aug 15;11(33):eads4473. doi: 10.1126/sciadv.ads4473. Epub 2025 Aug 15. Sci Adv. 2025. PMID: 40815652 Free PMC article.
Abstract
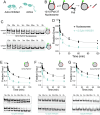
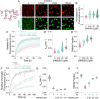
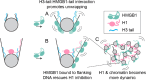
The essential architectural protein HMGB1 increases accessibility of nucleosomal DNA and counteracts the effects of linker histone H1. However, HMGB1 is less abundant than H1 and binds nucleosomes more weakly raising the question of how HMGB1 effectively competes with H1. Here, we show that HMGB1 rescues H1's inhibition of nucleosomal DNA accessibility without displacing H1. HMGB1 also increases the dynamics of condensed, H1-bound chromatin. Cryo-EM shows that HMGB1 binds at internal locations on a nucleosome and locally distorts the DNA. These sites, which are away from the binding site of H1, explain how HMGB1 and H1 co-occupy a nucleosome. Our findings lead to a model where HMGB1 counteracts the activity of H1 by distorting nucleosomal DNA and by contacting the H1 C-terminal tail. Compared to direct competition, nucleosome co-occupancy by HMGB1 and H1 allows a greater diversity of dynamic chromatin states and may be generalizable to other chromatin regulators.
Conflict of interest statement
Declaration of Interests Y.C. is scientific advisory board member of ShuiMu BioSciences. G.J.N. is a founder and scientific advisory board member of TippingPoint Biosciences.
Figures
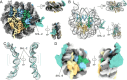



References
-
- Abdulhay N.J., Hsieh L.J., McNally C.P., Ostrowski M.S., Moore C.M., Ketavarapu M., Kasinathan S., Nanda A.S., Wu K., Chio U.S., Zhou Z., Goodarzi H., Narlikar G.J., Ramani V., 2023. Nucleosome density shapes kilobase-scale regulation by a mammalian chromatin remodeler. Nat Struct Mol Biol 30, 1571–1581. 10.1038/s41594-023-01093-6 - DOI - PMC - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
